

| Basic Information | |
|---|---|
| Species | Citrus clementina |
| Cazyme ID | Ciclev10007363m |
| Family | CBM43 |
| Protein Properties | Length: 959 Molecular Weight: 107186 Isoelectric Point: 5.4627 |
| Chromosome | Chromosome/Scaffold: 1 Start: 5280769 End: 5289488 |
| Description | O-Glycosyl hydrolases family 17 protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM43 | 849 | 930 | 5.8e-27 |
| WCVLNPHADDFEELPDSIDYACSLSDCTALGYGSSCNHLTVEGNASYAFNMYYQMNSQESWNCDFSGLAMVTDGDPSDDHCQ | |||
| GH17 | 506 | 826 | 0 |
| VGVNWGTMSTHHLPADNVVQMLRENRFNKLKLFEADKKILDALIGSDIEVMLAIPNYMLLEMSEDPGAAASWVYANVSSYFYTGGVNIKYVAVGNEPFLR TYNDTYRYITLPALKNIQQALNDAGLGSKIKATVPFNADIYNSPESNPVPSAGGFRDEVKDLTIGIIQFLHLNNAPFTVNIYPFLSLYGNDYFPVDFAFF EGTNKPIRDGSLLYTNVFDANFDTLVWSLDKAGYPDMHIIVGEVGWPTDGDKNANIENAKKFSQGLLKHVLSGAGTPARKGTIDVYLFSLLDENAKSIAP GNFERHWGIFEYDGKPKYELD | |||
| Full Sequence |
|---|
| Protein Sequence Length: 959 Download |
| MLDESKFDVH LKLWALRIPR EHCKVASKIL NGYLINRARV KPITEDPTCQ ENRYVILSEN 60 VQSSDLSDIP RQKLDELNKL CKIEVVSYSM TLGYSYWDAD YVLKQILPPG VEVPTSFETI 120 VNYQYRGHIA HLNIHDELLP FKDVIAKVIY DKNYPRIKTV VNKVGTIANE FRVPEFEILA 180 GEDNMVTEVK QYGATFKLDY SLVYWNSRLE HEHLRIISQF RPGETICDMF AGIGPFAIPA 240 AQKGCIVFAN DLNPDSVHYL KINAKVNKVD NYVRAYNMDA REFIRQLMTA PAGEINSESD 300 VFNLKACGNS GIQANKKTGI ENVGLDVQDK EVAGNITSNS EGLQNYCRNA DASVTATKRP 360 SDGCLEENGT TNSASGRKGK TSKRMKGSEL PNTKTWEHVD HIIMNLPASA LKFLDAFRGL 420 IQRQYWKGSL PWIHCYCFIR ANETEELIIS EAESALNACI QDPIFHKVRN VAPNKKMAQT 480 RALEYDHFLV LFLVFVMFEV SSGDGVGVNW GTMSTHHLPA DNVVQMLREN RFNKLKLFEA 540 DKKILDALIG SDIEVMLAIP NYMLLEMSED PGAAASWVYA NVSSYFYTGG VNIKYVAVGN 600 EPFLRTYNDT YRYITLPALK NIQQALNDAG LGSKIKATVP FNADIYNSPE SNPVPSAGGF 660 RDEVKDLTIG IIQFLHLNNA PFTVNIYPFL SLYGNDYFPV DFAFFEGTNK PIRDGSLLYT 720 NVFDANFDTL VWSLDKAGYP DMHIIVGEVG WPTDGDKNAN IENAKKFSQG LLKHVLSGAG 780 TPARKGTIDV YLFSLLDENA KSIAPGNFER HWGIFEYDGK PKYELDLSGK MKNKGLAPVE 840 GVAYMPRRWC VLNPHADDFE ELPDSIDYAC SLSDCTALGY GSSCNHLTVE GNASYAFNMY 900 YQMNSQESWN CDFSGLAMVT DGDPSDDHCQ FPVMIAHGSS LLLRGGILDT LLAVVGQC* 960 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG2520 | COG2520 | 5.0e-7 | 400 | 475 | 79 | + Predicted methyltransferase [General function prediction only] | ||
| COG2520 | COG2520 | 6.0e-61 | 112 | 290 | 180 | + Predicted methyltransferase [General function prediction only] | ||
| pfam02475 | Met_10 | 8.0e-64 | 115 | 296 | 190 | + Met-10+ like-protein. The methionine-10 mutant allele of N. crassa codes for a protein of unknown function. However, homologous proteins have been found in yeast suggesting this protein may be involved in methionine biosynthesis, transport and/or utilisation. | ||
| pfam00332 | Glyco_hydro_17 | 1.0e-76 | 506 | 826 | 323 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI28113.1 | 0 | 1 | 955 | 2 | 968 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002281488.1 | 0 | 1 | 477 | 2 | 478 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002281500.1 | 0 | 477 | 956 | 1 | 476 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002324967.1 | 0 | 506 | 943 | 9 | 447 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002515573.1 | 0 | 507 | 958 | 31 | 482 | Glucan endo-1,3-beta-glucosidase precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 506 | 827 | 1 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3ur8_B | 0 | 506 | 828 | 3 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3ur8_A | 0 | 506 | 828 | 3 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3ur7_B | 0 | 506 | 828 | 3 | 315 | A Chain A, Higher-density Crystal Structure Of Potato Endo-1,3-beta-glucanase |
| PDB | 3ur7_A | 0 | 506 | 828 | 3 | 315 | A Chain A, Higher-density Crystal Structure Of Potato Endo-1,3-beta-glucanase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
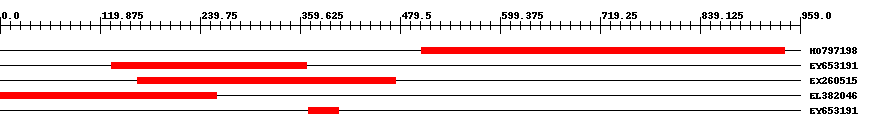 | ||||
| Hit | Length | Start | End | EValue |
| HO797198 | 437 | 505 | 940 | 0 |
| EY653191 | 234 | 134 | 367 | 0 |
| EX260515 | 310 | 165 | 474 | 0 |
| EL382046 | 260 | 1 | 260 | 0 |
| EY653191 | 37 | 370 | 406 | 0.000004 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |