

| Basic Information | |
|---|---|
| Species | Citrus clementina |
| Cazyme ID | Ciclev10027796m |
| Family | CBM43 |
| Protein Properties | Length: 864 Molecular Weight: 91214 Isoelectric Point: 7.8059 |
| Chromosome | Chromosome/Scaffold: 8 Start: 24795576 End: 24804627 |
| Description | O-Glycosyl hydrolases family 17 protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM43 | 368 | 447 | 2e-24 |
| WCVVNNNKDLSNATAIALEACSVADCSAISPGGSCYNISWPRNISYAFNSYYQQHNQRHDSCDFGRLGLITTVDPSADNC | |||
| GH17 | 24 | 348 | 0 |
| IGVNWGTNASHPLPPAKVVELLKANNIAKVKLFDADPLVLEALSGSNIGVTVGIPNSLLKSLNSSKKAAESWVHDNVTRYFSAGHGNGVRFEYVALGDEP FLQSYSEQFHPFIIGAAMNIQIALGRANLASEVKVVVPCSFDTFLSESGKPSGGHFRPDLNKTMIELLEFLSKHHSPFFVTISPFISFHQNKNISLDFAL FKETAHPHKDSHKTYKNSFDLSYDTLVTALSTVGFPEMDIVVAQIGWPTDGVANATSSTAEIFMKGLINHLRSRSGTPLRPRNPPIETYIYSLLDEDQRR TTTGNFERHWGVFTFDGQAKYHLNF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 864 Download |
| MFPYLTLLVL LLAIPSMGSL AGAIGVNWGT NASHPLPPAK VVELLKANNI AKVKLFDADP 60 LVLEALSGSN IGVTVGIPNS LLKSLNSSKK AAESWVHDNV TRYFSAGHGN GVRFEYVALG 120 DEPFLQSYSE QFHPFIIGAA MNIQIALGRA NLASEVKVVV PCSFDTFLSE SGKPSGGHFR 180 PDLNKTMIEL LEFLSKHHSP FFVTISPFIS FHQNKNISLD FALFKETAHP HKDSHKTYKN 240 SFDLSYDTLV TALSTVGFPE MDIVVAQIGW PTDGVANATS STAEIFMKGL INHLRSRSGT 300 PLRPRNPPIE TYIYSLLDED QRRTTTGNFE RHWGVFTFDG QAKYHLNFGQ NAEKLVNARN 360 VEYLPSRWCV VNNNKDLSNA TAIALEACSV ADCSAISPGG SCYNISWPRN ISYAFNSYYQ 420 QHNQRHDSCD FGRLGLITTV DPSADNCRFT IELRTSHSAS LHGAYLFLRT IGMSGMYGQE 480 GGDPAYVGGG YGGGGSAGVY GGYGGDGGGG YSGGGGRGGG YGGGRGGGGR GGGYGGGYSQ 540 NRGGDRGGRG GGGRGGGRGG SGRDGDWKCP NPSCGNLNFA RRVECNKCGA PSPAGASDRT 600 ASGGSGGYNR GGSGGGFGGN RGGRGDGGRA NYDGGRGTYE GRSGGGNRGS SYGGSQGRED 660 TSYGHVPPSS YPGASGNYPP ANSYGGNTNY EMDAVPPPTS YTGGPTSYPP SYGGPVGGYG 720 GDSFGDAGGG SRAGPPAAYD SGYAGGGPRY QGGSYSNAPA DTSAKVKQCD GNCDSYCDNS 780 RIYISNLPPD VTVDELRELF GGIGTVISHP FPLNQSFWFH VYFMSSLKRF CIYINAFFIM 840 HLSYWLNVCL SLLPVYHAIF WFL* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
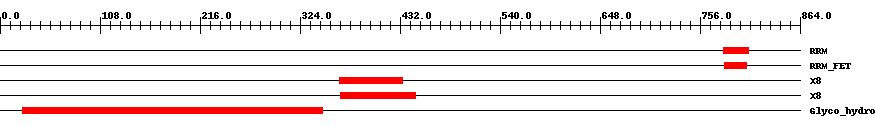 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| smart00360 | RRM | 1.0e-5 | 781 | 808 | 28 | + RNA recognition motif. | ||
| cd12280 | RRM_FET | 5.0e-6 | 782 | 806 | 25 | + RNA recognition motif in the FET family of RNA-binding proteins. This subfamily corresponds to the RRM of FET (previously TET) (FUS/TLS, EWS, TAF15) family of RNA-binding proteins. This ubiquitously expressed family of similarly structured proteins predominantly localizing to the nuclear, includes FUS (also known as TLS or Pigpen or hnRNP P2), EWS (also known as EWSR1), TAF15 (also known as hTAFII68 or TAF2N or RPB56), and Drosophila Cabeza (also known as SARFH). The corresponding coding genes of these proteins are involved in deleterious genomic rearrangements with transcription factor genes in a variety of human sarcomas and acute leukemias. All FET proteins interact with each other and are therefore likely to be part of the very same protein complexes, which suggests a general bridging role for FET proteins coupling RNA transcription, processing, transport, and DNA repair. The FET proteins contain multiple copies of a degenerate hexapeptide repeat motif at the N-terminus. The C-terminal region consists of a conserved nuclear import and retention signal (C-NLS), a putative zinc-finger domain, and a conserved RNA recognition motif (RRM), also known as RBD (RNA binding domain) or RNP (ribonucleoprotein domain), which is flanked by 3 arginine-glycine-glycine (RGG) boxes. FUS and EWS might have similar sequence specificity; both bind preferentially to GGUG-containing RNAs. FUS has also been shown to bind strongly to human telomeric RNA and to small low-copy-number RNAs tethered to the promoter of cyclin D1. To date, nothing is known about the RNA binding specificity of TAF15. | ||
| pfam07983 | X8 | 4.0e-14 | 367 | 435 | 79 | + X8 domain. The X8 domain domain contains at least 6 conserved cysteine residues that presumably form three disulphide bridges. The domain is found in an Olive pollen allergen as well as at the C-terminus of several families of glycosyl hydrolases. This domain may be involved in carbohydrate binding. This domain is characteristic of GPI-anchored domains. | ||
| smart00768 | X8 | 1.0e-27 | 368 | 449 | 84 | + Possibly involved in carbohydrate binding. The X8 domain, which may be involved in carbohydrate binding, is found in an Olive pollen antigen as well as at the C terminus of family 17 glycosyl hydrolases. It contains 6 conserved cysteine residues which presumably form three disulfide bridges. | ||
| pfam00332 | Glyco_hydro_17 | 2.0e-62 | 24 | 348 | 326 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003676 | nucleic acid binding |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005622 | intracellular |
| GO:0005975 | carbohydrate metabolic process |
| GO:0008270 | zinc ion binding |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI28434.1 | 0 | 21 | 471 | 20 | 471 | unnamed protein product [Vitis vinifera] |
| EMBL | CBI28434.1 | 0.00000000006 | 564 | 594 | 619 | 649 | unnamed protein product [Vitis vinifera] |
| EMBL | CBI28434.1 | 5e-16 | 690 | 806 | 693 | 758 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002269108.1 | 0 | 23 | 471 | 25 | 474 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002313970.1 | 0 | 23 | 474 | 21 | 471 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3f55_D | 0 | 24 | 349 | 2 | 316 | A Chain A, Structure Of S.Olivaceoviridis Xylanase Q88aR275A MUTANT |
| PDB | 3f55_C | 0 | 24 | 349 | 2 | 316 | A Chain A, Structure Of S.Olivaceoviridis Xylanase Q88aR275A MUTANT |
| PDB | 3f55_B | 0 | 24 | 349 | 2 | 316 | A Chain A, Structure Of S.Olivaceoviridis Xylanase Q88aR275A MUTANT |
| PDB | 3f55_A | 0 | 24 | 349 | 2 | 316 | A Chain A, Structure Of S.Olivaceoviridis Xylanase Q88aR275A MUTANT |
| PDB | 3em5_D | 0 | 24 | 349 | 2 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |