

| Basic Information | |
|---|---|
| Species | Prunus persica |
| Cazyme ID | ppa024453m |
| Family | GT1 |
| Protein Properties | Length: 451 Molecular Weight: 50211.9 Isoelectric Point: 5.1174 |
| Chromosome | Chromosome/Scaffold: 4 Start: 2638624 End: 2639976 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
| Family | Start | End | Evalue |
| GT1 | 219 | 412 | 2.3e-26 |
| DYVETQFGKPVILAGPVVPESPTTELDEKWAKWFDGFKAKTVIYCAFGSECILNKAQFQELLLGFELTGLPFFAALKPPMGVESIEAALPEGFEERVQGK GIVHGGWVQQPLILKHPSVGCFVTHCGSGSLSEALVNECRLVLLPNMGDQIINARMMSRDLKVGVEVEKEDEDGVFTTEGVCKAVKTVMDDESE | |||
| Full Sequence |
|---|
| Protein Sequence Length: 451 Download |
| MYPWFAMGHL TSFLHISNKL AQRGHRISFF SPSKVQQKLE AFNLHSHRIS FIPIDVPHVD 60 GLPPGTETTA DVPSLLQPLL VTAMDLTRPA IEQALCELRP NFVFFDFTHW LPELLHKLDI 120 GIKSVYYCTI SPATVGFLMC PERKLLEISL TEADLMEPPP SFPASSIKLQ THEARALVGA 180 SLMEFGRGVT FLERQMGSFS NCDAIAFKTC REMEGPYCDY VETQFGKPVI LAGPVVPESP 240 TTELDEKWAK WFDGFKAKTV IYCAFGSECI LNKAQFQELL LGFELTGLPF FAALKPPMGV 300 ESIEAALPEG FEERVQGKGI VHGGWVQQPL ILKHPSVGCF VTHCGSGSLS EALVNECRLV 360 LLPNMGDQII NARMMSRDLK VGVEVEKEDE DGVFTTEGVC KAVKTVMDDE SEVAKEVWTN 420 HAKWREFLSG KELENSCLDS FEQKLHQLPQ * |
| Functional Domains Download unfiltered results here | ||||||
|---|---|---|---|---|---|---|
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description |
| PLN02863 | PLN02863 | 7.0e-32 | 212 | 444 | 246 | + UDP-glucoronosyl/UDP-glucosyl transferase family protein |
| PLN02670 | PLN02670 | 1.0e-76 | 1 | 438 | 454 | + transferase, transferring glycosyl groups |
| PLN00414 | PLN00414 | 3.0e-135 | 1 | 445 | 450 | + glycosyltransferase family protein |
| PLN02208 | PLN02208 | 4.0e-140 | 1 | 447 | 449 | + glycosyltransferase family protein |
| PLN02764 | PLN02764 | 1.0e-145 | 1 | 448 | 450 | + glycosyltransferase family protein |
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016758 | transferase activity, transferring hexosyl groups |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABL74480.1 | 0 | 1 | 448 | 12 | 457 | glucosyltransferase [Ipomoea batatas] |
| GenBank | ACU24192.1 | 0 | 1 | 448 | 10 | 456 | unknown [Glycine max] |
| DDBJ | BAD95882.1 | 0 | 1 | 448 | 12 | 457 | glucosyltransferase [Ipomoea purpurea] |
| RefSeq | XP_002331944.1 | 0 | 1 | 448 | 1 | 444 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002522508.1 | 0 | 1 | 448 | 10 | 455 | UDP-glucosyltransferase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2vg8_A | 2e-20 | 228 | 410 | 235 | 430 | X Chain X, Structure Of Isoniazid (Inh) Bound To Cytosolic Soybean Ascorbate Peroxidase |
| PDB | 2vch_A | 2e-20 | 228 | 410 | 235 | 430 | X Chain X, Structure Of Isoniazid (Inh) Bound To Cytosolic Soybean Ascorbate Peroxidase |
| PDB | 2vce_A | 2e-20 | 228 | 410 | 235 | 430 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 2c9z_A | 1e-18 | 250 | 426 | 263 | 427 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| PDB | 2c1z_A | 1e-18 | 250 | 426 | 263 | 427 | A Chain A, Characterization And Engineering Of The Bifunctional N- And O-glucosyltransferase Involved In Xenobiotic Metabolism In Plants |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
| Hit | Length | Start | End | EValue |
| HO778286 | 441 | 12 | 448 | 0 |
| DY272554 | 328 | 1 | 328 | 0 |
| CB292845 | 301 | 10 | 310 | 0 |
| FC906269 | 280 | 1 | 280 | 0 |
| HO778286 | 23 | 1 | 23 | 0.004 |
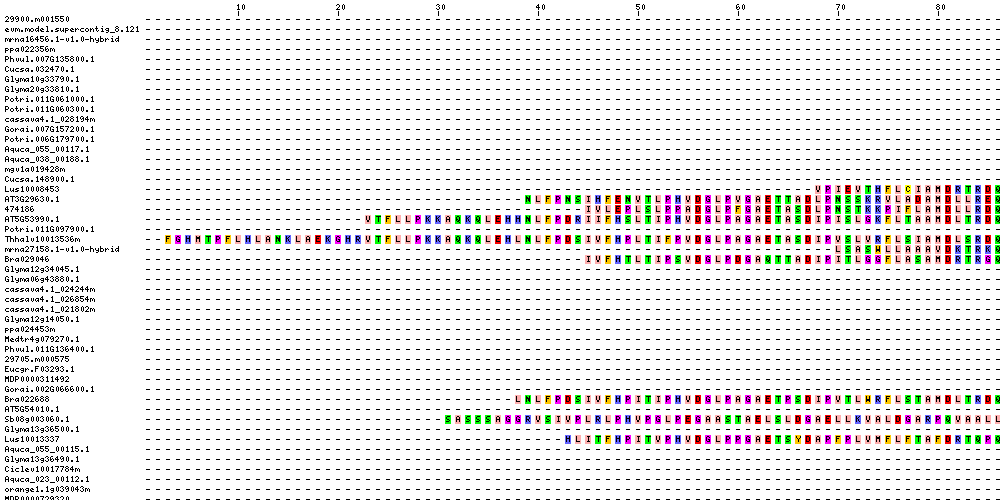
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |