

| Basic Information | |
|---|---|
| Species | Phaseolus vulgaris |
| Cazyme ID | Phvul.002G199300.1 |
| Family | AA7 |
| Protein Properties | Length: 574 Molecular Weight: 63341.6 Isoelectric Point: 8.8852 |
| Chromosome | Chromosome/Scaffold: 02 Start: 35777680 End: 35780225 |
| Description | FAD-binding Berberine family protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA7 | 71 | 532 | 0 |
| NARFNTTSTPKPLLVVTPLEEPHVQGAVICSKNIGIQLKIRSGGHDFEGISYVSDQPFIILDMFQFRNITVDVQNEIAVVQAGATLGEVYYRIWEKSKVH GFPAGVCPTVGVGGHLSGGGYGNMLRKHGLAVDHVIDAKIVDVKGRILDKKGMGEDLFWAIRGGGGASFGVILSYTIKLVPVPEVVTVYRVAKSLDKNES ATEFVLQWQQVAPHTDDRLFMRLLLQPVSSKVVKGQKTIRATVVALFLGGADEVVKLMGKEFPTLGLSKQNCTEMNWIDSVLWWANFDNITKPDALLDRN VNSASFLKRKSDYVQKPISKEGLEGIWKKMVELGKTGFVFNPYGGKMSEVSSDATPFPHRAGNLFKIQYSVNWDEPGVELENNFTRQARMLYSYMTPFVS SNPRSAFLNYRDLDIGTNSFGNNSYKEGAVYGVKYFNDNFDRLVKIKSAVDPENFFRNEQSI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 574 Download |
| MENLSLFPSG VLLFLLLSVT ASVAAPTAES VYTSFLQCLT NYTKSQDHVS DIVFAQTNSS 60 FSSVLQAYIR NARFNTTSTP KPLLVVTPLE EPHVQGAVIC SKNIGIQLKI RSGGHDFEGI 120 SYVSDQPFII LDMFQFRNIT VDVQNEIAVV QAGATLGEVY YRIWEKSKVH GFPAGVCPTV 180 GVGGHLSGGG YGNMLRKHGL AVDHVIDAKI VDVKGRILDK KGMGEDLFWA IRGGGGASFG 240 VILSYTIKLV PVPEVVTVYR VAKSLDKNES ATEFVLQWQQ VAPHTDDRLF MRLLLQPVSS 300 KVVKGQKTIR ATVVALFLGG ADEVVKLMGK EFPTLGLSKQ NCTEMNWIDS VLWWANFDNI 360 TKPDALLDRN VNSASFLKRK SDYVQKPISK EGLEGIWKKM VELGKTGFVF NPYGGKMSEV 420 SSDATPFPHR AGNLFKIQYS VNWDEPGVEL ENNFTRQARM LYSYMTPFVS SNPRSAFLNY 480 RDLDIGTNSF GNNSYKEGAV YGVKYFNDNF DRLVKIKSAV DPENFFRNEQ SIPTHTGLDA 540 GASKSGASKL SHFSLYWKLM AKVGGLLILE LLI* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
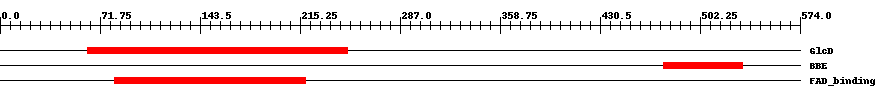
| COG0277 | GlcD | 3.0e-15 | 63 | 249 | 199 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| pfam08031 | BBE | 1.0e-15 | 476 | 533 | 58 | + Berberine and berberine like. This domain is found in the berberine bridge and berberine bridge- like enzymes which are involved in the biosynthesis of numerous isoquinoline alkaloids. They catalyze the transformation of the N-methyl group of (S)-reticuline into the C-8 berberine bridge carbon of (S)-scoulerine. | ||
| pfam01565 | FAD_binding_4 | 7.0e-18 | 82 | 219 | 140 | + FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidises the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_002270139.1 | 0 | 28 | 533 | 24 | 528 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002299028.1 | 0 | 11 | 534 | 12 | 531 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002317087.1 | 0 | 24 | 534 | 25 | 530 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002523157.1 | 0 | 30 | 534 | 35 | 534 | Reticuline oxidase precursor, putative [Ricinus communis] |
| RefSeq | XP_002523158.1 | 0 | 30 | 534 | 35 | 534 | Reticuline oxidase precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3vte_A | 0 | 34 | 533 | 6 | 511 | A Chain A, Crystal Structure Of Tetrahydrocannabinolic Acid Synthase From Cannabis Sativa |
| PDB | 4dns_B | 0 | 26 | 533 | 4 | 494 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 4dns_A | 0 | 26 | 533 | 4 | 494 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 3tsj_B | 0 | 35 | 533 | 11 | 494 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |
| PDB | 3tsj_A | 0 | 35 | 533 | 11 | 494 | A Chain A, Crystal Structure Of Bermuda Grass Isoallergen Bg60 Provides Insight Into The Various Cross-Allergenicity Of The Pollen Group 4 Allergens |