

| Basic Information | |
|---|---|
| Species | Glycine max |
| Cazyme ID | Glyma08g14730.1 |
| Family | AA1 |
| Protein Properties | Length: 577 Molecular Weight: 64852.6 Isoelectric Point: 8.0622 |
| Chromosome | Chromosome/Scaffold: 08 Start: 10724343 End: 10727358 |
| Description | Cupredoxin superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
| Family | Start | End | Evalue |
| AA1 | 349 | 552 | 4.48416e-44 |
| RLAQSFSIKARQGYIHKPPTTSDRVIVLLNTQNNISEYRHWSVNNVSFTLPHTPYLIALKENINGAFDSTPPPDGYDFANYDIFSVASNANATSSSGIYR LKFNTTVDIILQNANTMTKTNSETHPWHLHGHDFWVLGYGKGKFDVNNDTKKYNLENPIMKNTVPVHPFGWTALRFRTDNPGVWAFHCHIESHFYMGMGV VFEE | |||
| AA1 | 90 | 352 | 0 |
| LSIHWHGIRQIGTPWFDGTEGVTQCPILPGDTFIYQFVVDRPGTYLYHAHYGIQREAGLYGMMRVAPRDPEPFAYDLDRSIILNDWYHSSTYEQAAGLSS IPFRWVGEPQSLLIHGKGIFNCSKSPSLGTDVCDASKCSPFVQTVIPGKTYRLRIASLTALSALSFQIEGHNMTVVEADGHYVEPFVVKNLFIYSGETYS VTVKSDQDPSRNYWITSNVVSRNRSTPAGLGMFNYYPNHPKRSPPTVPPSPPAWHDVEPRLAQ | |||
| Full Sequence |
|---|
| Protein Sequence Length: 577 Download |
| MVELGLISLR ALQLPRLLIL CFFVILGNFH KAEARIRHYK WEAKYEFRSP DCFKKLVITI 60 NGKTPGPSIQ AQEGDTIIVQ VNNSLVTENL SIHWHGIRQI GTPWFDGTEG VTQCPILPGD 120 TFIYQFVVDR PGTYLYHAHY GIQREAGLYG MMRVAPRDPE PFAYDLDRSI ILNDWYHSST 180 YEQAAGLSSI PFRWVGEPQS LLIHGKGIFN CSKSPSLGTD VCDASKCSPF VQTVIPGKTY 240 RLRIASLTAL SALSFQIEGH NMTVVEADGH YVEPFVVKNL FIYSGETYSV TVKSDQDPSR 300 NYWITSNVVS RNRSTPAGLG MFNYYPNHPK RSPPTVPPSP PAWHDVEPRL AQSFSIKARQ 360 GYIHKPPTTS DRVIVLLNTQ NNISEYRHWS VNNVSFTLPH TPYLIALKEN INGAFDSTPP 420 PDGYDFANYD IFSVASNANA TSSSGIYRLK FNTTVDIILQ NANTMTKTNS ETHPWHLHGH 480 DFWVLGYGKG KFDVNNDTKK YNLENPIMKN TVPVHPFGWT ALRFRTDNPG VWAFHCHIES 540 HFYMGMGVVF EEGVERVGKL PSSIMGCGQT RGFHGP* |
| Functional Domains Download unfiltered results here | ||||||
|---|---|---|---|---|---|---|
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description |
| TIGR03390 | ascorbOXfungal | 5.0e-79 | 52 | 544 | 528 | + L-ascorbate oxidase, fungal type. This model describes a family of fungal ascorbate oxidases, within a larger family of multicopper oxidases that also includes plant ascorbate oxidases (TIGR03388), plant laccases and laccase-like proteins (TIGR03389), and related proteins. The member from Acremonium sp. HI-25 is characterized. |
| TIGR03389 | laccase | 4.0e-101 | 34 | 541 | 538 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. |
| PLN02604 | PLN02604 | 0 | 21 | 572 | 559 | + oxidoreductase |
| TIGR03388 | ascorbase | 0 | 36 | 570 | 543 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| PLN02191 | PLN02191 | 0 | 24 | 573 | 559 | + L-ascorbate oxidase |
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005507 | copper ion binding |
| GO:0016491 | oxidoreductase activity |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAY47050.1 | 0 | 32 | 576 | 31 | 578 | ascorbate oxidase [Solanum lycopersicum] |
| DDBJ | BAB86897.1 | 0 | 12 | 576 | 1 | 567 | syringolide-induced protein B13-1-1 [Glycine max] |
| RefSeq | XP_002281435.1 | 0 | 17 | 575 | 6 | 565 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002281497.1 | 0 | 12 | 575 | 1 | 565 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002306323.1 | 0 | 16 | 576 | 2 | 570 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1asq_B | 0 | 34 | 570 | 1 | 541 | A Chain A, Native Structure Of Beta-Galactosidase From Penicillium Sp. |
| PDB | 1asq_A | 0 | 34 | 570 | 1 | 541 | A Chain A, Native Structure Of Beta-Galactosidase From Penicillium Sp. |
| PDB | 1asp_B | 0 | 34 | 570 | 1 | 541 | A Chain A, Native Structure Of Beta-Galactosidase From Penicillium Sp. |
| PDB | 1asp_A | 0 | 34 | 570 | 1 | 541 | A Chain A, Native Structure Of Beta-Galactosidase From Penicillium Sp. |
| PDB | 1aso_B | 0 | 34 | 570 | 1 | 541 | A Chain A, Native Structure Of Beta-Galactosidase From Penicillium Sp. |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
| Hit | Length | Start | End | EValue |
| GR847295 | 260 | 71 | 328 | 0 |
| EL420184 | 276 | 32 | 303 | 0 |
| DV705515 | 300 | 245 | 544 | 0 |
| CK298807 | 284 | 23 | 298 | 0 |
| DV706195 | 292 | 245 | 536 | 0 |
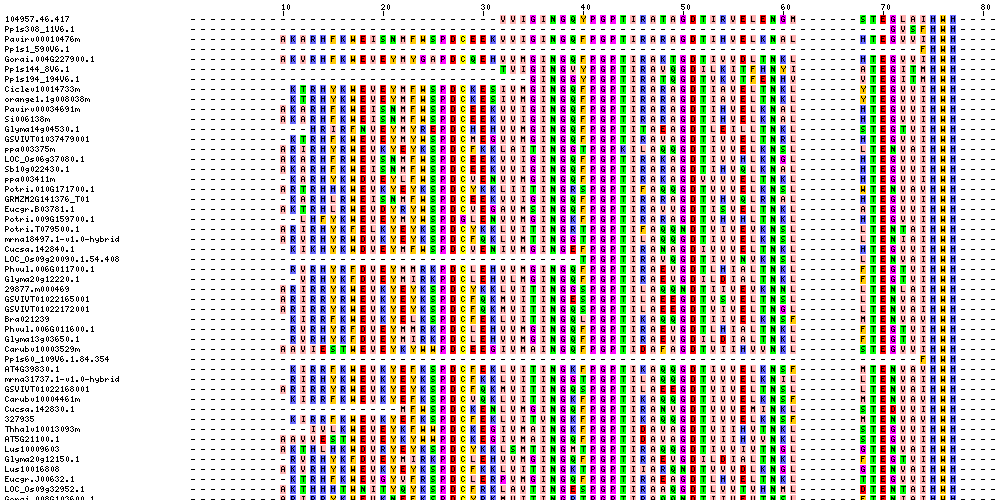
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |