

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s115_92V6.1 |
| Family | GT47 |
| Protein Properties | Length: 396 Molecular Weight: 44776.8 Isoelectric Point: 8.2848 |
| Chromosome | Chromosome/Scaffold: 115 Start: 602134 End: 603321 |
| Description | Exostosin family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT47 | 129 | 342 | 1e-32 |
| TLPYWNGGLNHVLVIFADKWHQKGPHQASIGNASVMASDIHETTYRPGFDVSIPLPGNHHLREFQSVKPLERKYLATFRGLRYLGLKGEGVFRSLESFRG MHNGNDVIVATSCEHPISKMIRKKEPALGVHCDEDLLIHKNYTFNDLMNTTFGLVPAGVQPASYRFIEVLSAGAIPVLIADNYVKPFDTLILWYKCLLQF PTTEMHRIVGTLRA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 396 Download |
| MKFLASTPAN CTVCAVCPPA ETVVVEKEIV KEVFKKAECG NKVEPWYAEG KMWHYPARFP 60 LCSMDVCFNY SKCDGSEDLL IFTYNLPAPP KRYFSRINES KYYTNDPAKA CLFLVFLDGD 120 NPWPPHPSTL PYWNGGLNHV LVIFADKWHQ KGPHQASIGN ASVMASDIHE TTYRPGFDVS 180 IPLPGNHHLR EFQSVKPLER KYLATFRGLR YLGLKGEGVF RSLESFRGMH NGNDVIVATS 240 CEHPISKMIR KKEPALGVHC DEDLLIHKNY TFNDLMNTTF GLVPAGVQPA SYRFIEVLSA 300 GAIPVLIADN YVKPFDTLIL WYKCLLQFPT TEMHRIVGTL RAMKPEEIKT RQENCLAIYN 360 KYLKDDETLL RTTIQALKTR FLGAIPGFTD ITGKR* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam03016 | Exostosin | 6.0e-34 | 94 | 343 | 274 | + Exostosin family. The EXT family is a family of tumour suppressor genes. Mutations of EXT1 on 8q24.1, EXT2 on 11p11-13, and EXT3 on 19p have been associated with the autosomal dominant disorder known as hereditary multiple exostoses (HME). This is the most common known skeletal dysplasia. The chromosomal locations of other EXT genes suggest association with other forms of neoplasia. EXT1 and EXT2 have both been shown to encode a heparan sulphate polymerase with both D-glucuronyl (GlcA) and N-acetyl-D-glucosaminoglycan (GlcNAC) transferase activities. The nature of the defect in heparan sulphate biosynthesis in HME is unclear. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0016020 | membrane |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_001754325.1 | 0 | 62 | 369 | 1 | 308 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001761172.1 | 0 | 52 | 388 | 1 | 343 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001761253.1 | 0 | 62 | 394 | 1 | 336 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001769488.1 | 0 | 52 | 395 | 1 | 344 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001771909.1 | 0 | 62 | 369 | 1 | 308 | predicted protein [Physcomitrella patens subsp. patens] |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
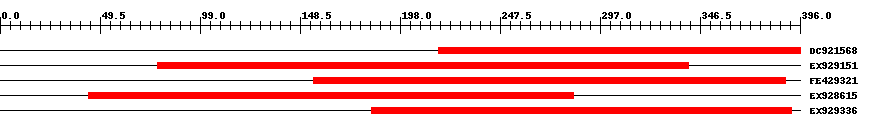 | ||||
| Hit | Length | Start | End | EValue |
| DC921568 | 180 | 217 | 396 | 0 |
| EX929151 | 265 | 78 | 341 | 0 |
| FE429321 | 236 | 155 | 389 | 0 |
| EX928615 | 242 | 44 | 284 | 0 |
| EX929336 | 209 | 184 | 392 | 0 |
| Orthologous Group | |||||
|---|---|---|---|---|---|
| Species | ID | ||||
| Physcomitrella patens | Pp1s143_125V6.1 | Pp1s47_67V6.1 | Pp1s13_257V6.1 | Pp1s47_134V6.1 | Pp1s47_145V6.1 |
| Pp1s69_76V6.1 | Pp1s68_175V6.1 | Pp1s111_147V6.1 | |||
| Selaginella moellendorffii | 23945 | ||||
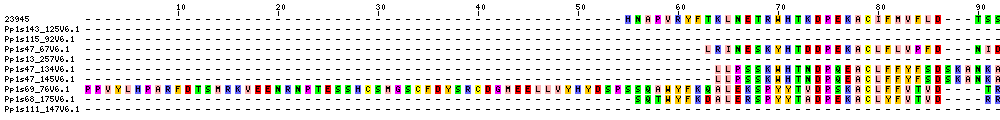
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |