

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s13_257V6.1 |
| Family | GT47 |
| Protein Properties | Length: 444 Molecular Weight: 50386.2 Isoelectric Point: 8.4147 |
| Chromosome | Chromosome/Scaffold: 13 Start: 1717872 End: 1719203 |
| Description | Exostosin family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT47 | 177 | 390 | 1.8e-32 |
| TLPYWNGGQNHVLVIFADKWHQKGPHQASIGNASVMASDTHETTYRPGFDVSVPLPGRLHVREFQSLKPLERKYLATFRGLRYLGLKGEGVFRSLDAFRG MHNGNDVIVATSCEHAISKMIREKEPALGVHCDEDLLIHKNFTFHDLMNTTFGLVPAGVQPASYRFIEVLSAGAIPVLIADNYVKPFDTLILWYKCLLQF PTTEMHRIVGTLRA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 444 Download |
| MKRVVCRLEL ILFLCFVVIF VSLTSFNSAK FLKHYNSIVS VSRDPGTKLL ESTPANCNCT 60 DRAVCPPAET VVVEKEIIKE VVKKEECRNN GGPWYAEGKT WHYPARFPLC SMDVCFNYTK 120 CDSSENLLIF TYNLPSPPKR YFSRINESKY YTNDPAKACL FLVFLDGDTP WPPHPSTLPY 180 WNGGQNHVLV IFADKWHQKG PHQASIGNAS VMASDTHETT YRPGFDVSVP LPGRLHVREF 240 QSLKPLERKY LATFRGLRYL GLKGEGVFRS LDAFRGMHNG NDVIVATSCE HAISKMIREK 300 EPALGVHCDE DLLIHKNFTF HDLMNTTFGL VPAGVQPASY RFIEVLSAGA IPVLIADNYV 360 KPFDTLILWY KCLLQFPTTE MHRIVGTLRA MKPEEIQTRQ ENCLAIYNKY LKDDETLLQT 420 SIQALKIRFL GTIPGFTDIT RKR* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam03016 | Exostosin | 1.0e-32 | 123 | 391 | 314 | + Exostosin family. The EXT family is a family of tumour suppressor genes. Mutations of EXT1 on 8q24.1, EXT2 on 11p11-13, and EXT3 on 19p have been associated with the autosomal dominant disorder known as hereditary multiple exostoses (HME). This is the most common known skeletal dysplasia. The chromosomal locations of other EXT genes suggest association with other forms of neoplasia. EXT1 and EXT2 have both been shown to encode a heparan sulphate polymerase with both D-glucuronyl (GlcA) and N-acetyl-D-glucosaminoglycan (GlcNAC) transferase activities. The nature of the defect in heparan sulphate biosynthesis in HME is unclear. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0016020 | membrane |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | XP_001754325.1 | 0 | 110 | 417 | 1 | 308 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001761172.1 | 0 | 101 | 436 | 2 | 343 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001761253.1 | 0 | 110 | 442 | 1 | 336 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001769488.1 | 0 | 101 | 443 | 2 | 344 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001771909.1 | 0 | 110 | 417 | 1 | 308 | predicted protein [Physcomitrella patens subsp. patens] |
| Transmembrane Domains | |||||
|---|---|---|---|---|---|
 | |||||
| Start | End | ||||
| 7 | 29 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DC921568 | 180 | 265 | 444 | 0 |
| DC907675 | 182 | 1 | 182 | 0 |
| EX929151 | 265 | 126 | 389 | 0 |
| FE429321 | 236 | 203 | 437 | 0 |
| EX928615 | 243 | 91 | 332 | 0 |
| Orthologous Group | |||||
|---|---|---|---|---|---|
| Species | ID | ||||
| Physcomitrella patens | Pp1s143_125V6.1 | Pp1s115_92V6.1 | Pp1s47_67V6.1 | Pp1s47_134V6.1 | Pp1s47_145V6.1 |
| Pp1s69_76V6.1 | Pp1s68_175V6.1 | Pp1s111_147V6.1 | |||
| Selaginella moellendorffii | 23945 | ||||
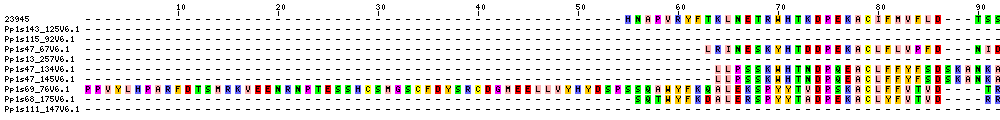
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |