

| Basic Information | |
|---|---|
| Species | Capsella rubella |
| Cazyme ID | Carubv10019007m |
| Family | GH28 |
| Protein Properties | Length: 478 Molecular Weight: 52227.6 Isoelectric Point: 4.8162 |
| Chromosome | Chromosome/Scaffold: 5 Start: 11686734 End: 11688416 |
| Description | Pectin lyase-like superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH28 | 74 | 399 | 0 |
| SICRVVFPSGTYLTAKLHLRSGVVLDVTENAVILGGPRIEDYYPADTSSDWYVVVANNATDVGITGGGAIDGQGSKFVVRFDEKKNIMVSWNQTGACLGD ECRPRLVGFVDSTNVEIWNITLREPPYWCLHIVRCENTSVHDVSILGDFNTPNNDGIDIEDSNNTVITRCHIDTGDDAICPKTYTGPLYNLTATDCWIRT KSSAIKLGSASWFDFKGLFFDNITIFESHRGLGMQIRDGGNVSDITFSNINISTRYYDPSWWGRAEPIYITSCPRSPSAKEGAISNLLFVNITINSENGV FLSGSPNGLLSDIKFKNMNLTVRRWS | |||
| Full Sequence |
|---|
| Protein Sequence Length: 478 Download |
| MGTSLTGSIL LLLLFLSLVQ SHSYTSFSKI QLPGDSLTLS VTDFGATGDG ISYDSSAIQS 60 TIDACNRHYA SSSSICRVVF PSGTYLTAKL HLRSGVVLDV TENAVILGGP RIEDYYPADT 120 SSDWYVVVAN NATDVGITGG GAIDGQGSKF VVRFDEKKNI MVSWNQTGAC LGDECRPRLV 180 GFVDSTNVEI WNITLREPPY WCLHIVRCEN TSVHDVSILG DFNTPNNDGI DIEDSNNTVI 240 TRCHIDTGDD AICPKTYTGP LYNLTATDCW IRTKSSAIKL GSASWFDFKG LFFDNITIFE 300 SHRGLGMQIR DGGNVSDITF SNINISTRYY DPSWWGRAEP IYITSCPRSP SAKEGAISNL 360 LFVNITINSE NGVFLSGSPN GLLSDIKFKN MNLTVRRWSN YSAGLVDYRP GCQGLVNHST 420 TAGIIMEHVN GFSVENVDLK WSDDLNSAWN VPLEFRPSTV NNISFVGFNS GLLHEIV* 480 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
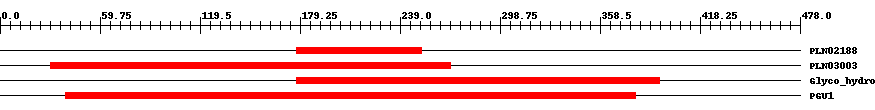 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02188 | PLN02188 | 6.0e-8 | 177 | 252 | 76 | + polygalacturonase/glycoside hydrolase family protein | ||
| PLN03003 | PLN03003 | 1.0e-8 | 30 | 269 | 243 | + Probable polygalacturonase At3g15720 | ||
| pfam00295 | Glyco_hydro_28 | 2.0e-12 | 177 | 394 | 233 | + Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes is important in cell wall metabolism. | ||
| COG5434 | PGU1 | 1.0e-40 | 39 | 380 | 388 | + Endopygalactorunase [Cell envelope biogenesis, outer membrane] | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004650 | polygalacturonase activity |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAB41176.1 | 0 | 5 | 476 | 1 | 474 | putative protein [Arabidopsis thaliana] |
| RefSeq | NP_567055.1 | 0 | 5 | 476 | 1 | 474 | glycoside hydrolase family 28 protein / polygalacturonase (pectinase) family protein [Arabidopsis thaliana] |
| RefSeq | XP_002282920.1 | 0 | 9 | 471 | 8 | 465 | PREDICTED: hypothetical protein isoform 1 [Vitis vinifera] |
| RefSeq | XP_002282925.1 | 0 | 9 | 471 | 8 | 460 | PREDICTED: hypothetical protein isoform 2 [Vitis vinifera] |
| RefSeq | XP_002322693.1 | 0 | 28 | 476 | 1 | 445 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3jur_D | 3e-18 | 31 | 256 | 22 | 267 | A Chain A, The Crystal Structure Of A Hyperthermoactive Exopolygalacturonase From Thermotoga Maritima |
| PDB | 3jur_C | 3e-18 | 31 | 256 | 22 | 267 | A Chain A, The Crystal Structure Of A Hyperthermoactive Exopolygalacturonase From Thermotoga Maritima |
| PDB | 3jur_B | 3e-18 | 31 | 256 | 22 | 267 | A Chain A, The Crystal Structure Of A Hyperthermoactive Exopolygalacturonase From Thermotoga Maritima |
| PDB | 3jur_A | 3e-18 | 31 | 256 | 22 | 267 | A Chain A, The Crystal Structure Of A Hyperthermoactive Exopolygalacturonase From Thermotoga Maritima |
| PDB | 2uvf_B | 0.000000000000002 | 39 | 322 | 157 | 484 | A Chain A, The Crystal Structure Of A Hyperthermoactive Exopolygalacturonase From Thermotoga Maritima |