

| Basic Information | |
|---|---|
| Species | Phaseolus vulgaris |
| Cazyme ID | Phvul.009G144700.1 |
| Family | PL4 |
| Protein Properties | Length: 665 Molecular Weight: 76315.4 Isoelectric Point: 5.5518 |
| Chromosome | Chromosome/Scaffold: 09 Start: 21135575 End: 21140251 |
| Description | Rhamnogalacturonate lyase family protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL4 | 28 | 648 | 0 |
| RLDIQDKHVIMDNGIVQVNLSNPGGIVTGIQYNGIDNLLEVLNEEYDRGYWDIVWDQGGENRTKKGAKGKGTFDRMEATNLTVIVENEEQVELSFTRTWN VSFVGKLPPLNIDKRFIMLRGSSGFYTYAIYEHLKEWPAFDLDNTRIAFKPRKEKFHYMSVADNRQRFMPLPDDRIPPRGQVLAYPEAVRLVDPVEPEFK GEVDDKYEYSCESRYNQVHGWMSIDPLNSTGFWLITPSYEFRSAGPLKQYLTSHVGPTTLSVFHSTHYSGTDLIMQFGPNEPWKKVYGPIFIYLNSLSNG VSPIKLWEDAKQQMVSEVESWPYTFPASKDFLSSEERGKVQGRLLVRDRIVSDAFIPVSGAYVGLAAIGDVGSWQRECKGYQFWTITDDKGYFSIINVRP GDYNLYSWVTGYIGDYKFDKIINVTSGCEINVSELVYEPPRDGPTLWEIGIPDRSAAEFYVPDPNPMYINKLYVNHTDRFRQYGLWERYADLYPNKDLVY TVGVSDYKKDWFFAQVNRRKDNGTYQGTTWQINFNLDRVNTNGSYTLRVALASVHGAELQIRINKLEADPALFSSGVIGKENTIARHGIHGFYWLFSIDV DGTLLVQGNNTIFLTQPMNTS | |||
| Full Sequence |
|---|
| Protein Sequence Length: 665 Download |
| MDFTYHFLFL MMLQFLVALG ADTKPGVRLD IQDKHVIMDN GIVQVNLSNP GGIVTGIQYN 60 GIDNLLEVLN EEYDRGYWDI VWDQGGENRT KKGAKGKGTF DRMEATNLTV IVENEEQVEL 120 SFTRTWNVSF VGKLPPLNID KRFIMLRGSS GFYTYAIYEH LKEWPAFDLD NTRIAFKPRK 180 EKFHYMSVAD NRQRFMPLPD DRIPPRGQVL AYPEAVRLVD PVEPEFKGEV DDKYEYSCES 240 RYNQVHGWMS IDPLNSTGFW LITPSYEFRS AGPLKQYLTS HVGPTTLSVF HSTHYSGTDL 300 IMQFGPNEPW KKVYGPIFIY LNSLSNGVSP IKLWEDAKQQ MVSEVESWPY TFPASKDFLS 360 SEERGKVQGR LLVRDRIVSD AFIPVSGAYV GLAAIGDVGS WQRECKGYQF WTITDDKGYF 420 SIINVRPGDY NLYSWVTGYI GDYKFDKIIN VTSGCEINVS ELVYEPPRDG PTLWEIGIPD 480 RSAAEFYVPD PNPMYINKLY VNHTDRFRQY GLWERYADLY PNKDLVYTVG VSDYKKDWFF 540 AQVNRRKDNG TYQGTTWQIN FNLDRVNTNG SYTLRVALAS VHGAELQIRI NKLEADPALF 600 SSGVIGKENT IARHGIHGFY WLFSIDVDGT LLVQGNNTIF LTQPMNTSPF IGIMYDYIRL 660 EYKR* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
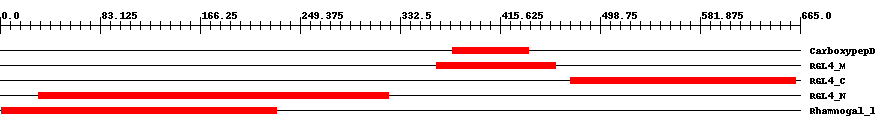 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam13620 | CarboxypepD_reg | 0.005 | 376 | 439 | 64 | + Carboxypeptidase regulatory-like domain. | ||
| cd10316 | RGL4_M | 2.0e-26 | 363 | 462 | 100 | + Middle domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold. Both the middle domain represented by this model and the C-terminal domain are putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. | ||
| cd10317 | RGL4_C | 8.0e-54 | 474 | 661 | 190 | + C-terminal domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold. Both the middle and the C-terminal domain are putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. | ||
| cd10320 | RGL4_N | 4.0e-65 | 32 | 323 | 294 | + N-terminal catalytic domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold; the middle and C-terminal domains are both putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. | ||
| pfam06045 | Rhamnogal_lyase | 2.0e-95 | 1 | 230 | 230 | + Rhamnogalacturonate lyase family. Rhamnogalacturonate lyase (EC:4.2.2.-) degrades the rhamnogalacturonan I (RG-I) backbone of pectin. This family contains mainly members from plants, but also contains the plant pathogen Erwinia chrysanthemi. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI19298.1 | 0 | 26 | 661 | 5 | 642 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002285626.1 | 0 | 38 | 661 | 1 | 613 | PREDICTED: hypothetical protein isoform 1 [Vitis vinifera] |
| RefSeq | XP_002527353.1 | 0 | 26 | 661 | 5 | 631 | lyase, putative [Ricinus communis] |
| RefSeq | XP_002527356.1 | 0 | 35 | 661 | 127 | 744 | lyase, putative [Ricinus communis] |
| RefSeq | XP_002527356.1 | 0 | 103 | 230 | 1 | 128 | lyase, putative [Ricinus communis] |