

| Basic Information | |
|---|---|
| Species | Setaria italica |
| Cazyme ID | Si012557m |
| Family | PL4 |
| Protein Properties | Length: 686 Molecular Weight: 75781.6 Isoelectric Point: 7.548 |
| Chromosome | Chromosome/Scaffold: 7 Start: 34680576 End: 34684616 |
| Description | Rhamnogalacturonate lyase family protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL4 | 40 | 656 | 0 |
| AAAHGVTLRVDHHQVLVDNGVVQVTLSKPQGHITAVRYNGERSLLHFAGQENTGGYWDVVWNYPGSDHPRGMIDMLDSTEFKVVSSSPEQVELSFRSIYN PSRQDSVRLNVDKRIVVLKGSSGFYCYAIFEHASNWPAMNISEARLAFKLNTDKFNYMAISDDIQRYMPSAADRDEPRGTALAYKEAVLLVNPQEPQFKG EVDDKYQYSLDNKDNVVHGWISSNHPSPMGFWVITPSNEFKSGGPMKRELTSHVGPTSLTMFLGTHYIGDDIVLNVGNGEYWKKVLGPVFIYLNSSPKHG DLRALWQDAKAQAQTEVSKWPYSFPKSPDFAKAGERGSVTGRLMVRDRFMRNDDMPAGMAYIGLAAPGQPGSWATECKGYQFWTTATSCGSFTIGNVRAG VYNLYAWVPGVLGDYMYTSAVTVTPGCAIDLGDLVFLPPRSGPTLWEIGVPDRTAAEFFIPDVDPRYANRLFLHRDMYRQYGLWERYAELYPDSDPVFTV GRSNHSKDWFFAHVTRKVGNGNVPTTRQIRFNLDHVVADGTYTLRIALAAAQMSRLQVHVNGGGMRRGGGVFTTPEFGGGNAIARHGIHGVQWSFEFPIR GYLLEEGENRISITQTR | |||
| Full Sequence |
|---|
| Protein Sequence Length: 686 Download |
| MAASAAGATR SPLLLPVILL VAAVGASSSL AAAAAALPNA AAHGVTLRVD HHQVLVDNGV 60 VQVTLSKPQG HITAVRYNGE RSLLHFAGQE NTGGYWDVVW NYPGSDHPRG MIDMLDSTEF 120 KVVSSSPEQV ELSFRSIYNP SRQDSVRLNV DKRIVVLKGS SGFYCYAIFE HASNWPAMNI 180 SEARLAFKLN TDKFNYMAIS DDIQRYMPSA ADRDEPRGTA LAYKEAVLLV NPQEPQFKGE 240 VDDKYQYSLD NKDNVVHGWI SSNHPSPMGF WVITPSNEFK SGGPMKRELT SHVGPTSLTM 300 FLGTHYIGDD IVLNVGNGEY WKKVLGPVFI YLNSSPKHGD LRALWQDAKA QAQTEVSKWP 360 YSFPKSPDFA KAGERGSVTG RLMVRDRFMR NDDMPAGMAY IGLAAPGQPG SWATECKGYQ 420 FWTTATSCGS FTIGNVRAGV YNLYAWVPGV LGDYMYTSAV TVTPGCAIDL GDLVFLPPRS 480 GPTLWEIGVP DRTAAEFFIP DVDPRYANRL FLHRDMYRQY GLWERYAELY PDSDPVFTVG 540 RSNHSKDWFF AHVTRKVGNG NVPTTRQIRF NLDHVVADGT YTLRIALAAA QMSRLQVHVN 600 GGGMRRGGGV FTTPEFGGGN AIARHGIHGV QWSFEFPIRG YLLEEGENRI SITQTRAFGE 660 FLGVMYDYIR LEGPPGSWRD PTRRA* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
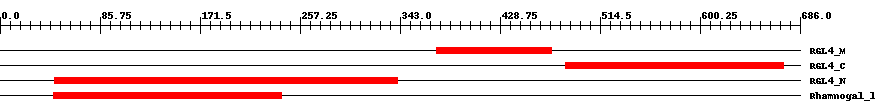 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| cd10316 | RGL4_M | 1.0e-30 | 374 | 473 | 100 | + Middle domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold. Both the middle domain represented by this model and the C-terminal domain are putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. | ||
| cd10317 | RGL4_C | 9.0e-48 | 485 | 672 | 190 | + C-terminal domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold. Both the middle and the C-terminal domain are putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. | ||
| cd10320 | RGL4_N | 6.0e-66 | 47 | 341 | 298 | + N-terminal catalytic domain of rhamnogalacturonan lyase, a family 4 polysaccharide lyase. The rhamnogalacturonan lyase of the polysaccharide lyase family 4 (RGL4) is involved in the degradation of RG (rhamnogalacturonan) type-I, an important pectic plant cell wall polysaccharide, by cleaving the alpha-1,4 glycoside bond between L-rhamnose and D-galacturonic acids in the backbone of RG type-I through a beta-elimination reaction. RGL4 consists of three domains, an N-terminal catalytic domain, a middle domain with a FNIII type fold and a C-terminal domain with a jelly roll fold; the middle and C-terminal domains are both putative carbohydrate binding modules. There are two types of RG lyases, which both cleave the alpha-1,4 bonds of the RG-I main chain (RG chain) through the beta-elimination reaction, but belong to two structurally unrelated polysaccharide lyase (PL) families, 4 and 11. | ||
| pfam06045 | Rhamnogal_lyase | 2.0e-95 | 46 | 241 | 196 | + Rhamnogalacturonate lyase family. Rhamnogalacturonate lyase (EC:4.2.2.-) degrades the rhamnogalacturonan I (RG-I) backbone of pectin. This family contains mainly members from plants, but also contains the plant pathogen Erwinia chrysanthemi. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABA96404.1 | 0 | 3 | 681 | 2 | 696 | LG27/30-like gene, putative, expressed [Oryza sativa (japonica cultivar-group)] |
| DDBJ | BAF29096.2 | 0 | 3 | 674 | 2 | 689 | Os12g0131900 [Oryza sativa Japonica Group] |
| GenBank | EEC68814.1 | 0 | 31 | 681 | 34 | 653 | hypothetical protein OsI_37377 [Oryza sativa Indica Group] |
| RefSeq | NP_001066077.1 | 0 | 3 | 681 | 2 | 683 | Os12g0131900 [Oryza sativa (japonica cultivar-group)] |
| RefSeq | XP_002441731.1 | 0 | 1 | 681 | 1 | 719 | hypothetical protein SORBIDRAFT_08g001440 [Sorghum bicolor] |