

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_10430876g0010 |
| Family | AA1 |
| Protein Properties | Length: 376 Molecular Weight: 41441.5 Isoelectric Point: 7.1606 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
| Family | Start | End | Evalue |
| AA1 | 117 | 362 | 0 |
| YSSAQSLNLPIIMADLPAMRDTNYATKFATGLRSLNSREYPSDIAKKIDRRLFLAVSLNLQDCPANSICKGYFGGRFSASINNQSFVRPKTALLQAAYYN ITGIFSKDFPSRPFHEYNYTDPILAVPNMNPNFSTRVAVIPYNANVELILQGTSIMGFENHPVHIHGFNVYVVGQGFGNYDPNTDPASFNLVDPPLRNTV GVPTAGWVALRFKADNPGIWFMHCHLELHTTWGLAMAFLVENGTGP | |||
| Full Sequence |
|---|
| Protein Sequence Length: 376 Download |
| MSLLMTADQP VGKYIMAARP YVTAGVPFDE TTATAILEYS SAQSLKLPII MADLPAMRDT 60 YNATKFATGL RSLNSREYPS DVPKKIDRRL FLAVSLNLQD CPAXXXXETT ATAILEYSSA 120 QSLNLPIIMA DLPAMRDTNY ATKFATGLRS LNSREYPSDI AKKIDRRLFL AVSLNLQDCP 180 ANSICKGYFG GRFSASINNQ SFVRPKTALL QAAYYNITGI FSKDFPSRPF HEYNYTDPIL 240 AVPNMNPNFS TRVAVIPYNA NVELILQGTS IMGFENHPVH IHGFNVYVVG QGFGNYDPNT 300 DPASFNLVDP PLRNTVGVPT AGWVALRFKA DNPGIWFMHC HLELHTTWGL AMAFLVENGT 360 GPAQSILPLP PDLPPC |
| Functional Domains Download unfiltered results here | ||||||
|---|---|---|---|---|---|---|
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description |
| TIGR03389 | laccase | 1.0e-27 | 1 | 102 | 102 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. |
| PLN02604 | PLN02604 | 9.0e-37 | 191 | 354 | 168 | + oxidoreductase |
| pfam07731 | Cu-oxidase_2 | 4.0e-41 | 234 | 359 | 126 | + Multicopper oxidase. This entry contains many divergent copper oxidase-like domains that are not recognised by the pfam00394 model. |
| TIGR03388 | ascorbase | 4.0e-42 | 109 | 354 | 251 | + L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| TIGR03389 | laccase | 1.0e-126 | 109 | 376 | 268 | + laccase, plant. Members of this protein family include the copper-containing enzyme laccase (EC 1.10.3.2), often several from a single plant species, and additional, uncharacterized, closely related plant proteins termed laccase-like multicopper oxidases. This protein family shows considerable sequence similarity to the L-ascorbate oxidase (EC 1.10.3.3) family. Laccases are enzymes of rather broad specificity, and classification of all proteins scoring about the trusted cutoff of this model as laccases may be appropriate. |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACN39933.1 | 0.00000000000002 | 4 | 114 | 272 | 382 | unknown [Picea sitchensis] |
| GenBank | ACN39933.1 | 0 | 89 | 376 | 281 | 559 | unknown [Picea sitchensis] |
| EMBL | CAN74557.1 | 6e-16 | 2 | 103 | 266 | 380 | hypothetical protein [Vitis vinifera] |
| EMBL | CAN74557.1 | 0 | 109 | 376 | 298 | 576 | hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002281603.1 | 0 | 109 | 376 | 305 | 583 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1asq_B | 9e-24 | 163 | 354 | 342 | 521 | A Chain A, Heme Ligand Mutant Of Recombinant Horseradish Peroxidase In Complex With Benzhydroxamic Acid |
| PDB | 1asq_A | 9e-24 | 163 | 354 | 342 | 521 | A Chain A, Heme Ligand Mutant Of Recombinant Horseradish Peroxidase In Complex With Benzhydroxamic Acid |
| PDB | 1asp_B | 9e-24 | 163 | 354 | 342 | 521 | A Chain A, Heme Ligand Mutant Of Recombinant Horseradish Peroxidase In Complex With Benzhydroxamic Acid |
| PDB | 1asp_A | 9e-24 | 163 | 354 | 342 | 521 | A Chain A, Heme Ligand Mutant Of Recombinant Horseradish Peroxidase In Complex With Benzhydroxamic Acid |
| PDB | 1aso_B | 9e-24 | 163 | 354 | 342 | 521 | A Chain A, Heme Ligand Mutant Of Recombinant Horseradish Peroxidase In Complex With Benzhydroxamic Acid |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
| Hit | Length | Start | End | EValue |
| DR465192 | 242 | 135 | 376 | 0 |
| FE520677 | 270 | 110 | 376 | 0 |
| FD730574 | 192 | 185 | 376 | 0 |
| DR465192 | 46 | 57 | 102 | 0.000000000002 |
| FE520677 | 75 | 29 | 103 | 0.000000001 |
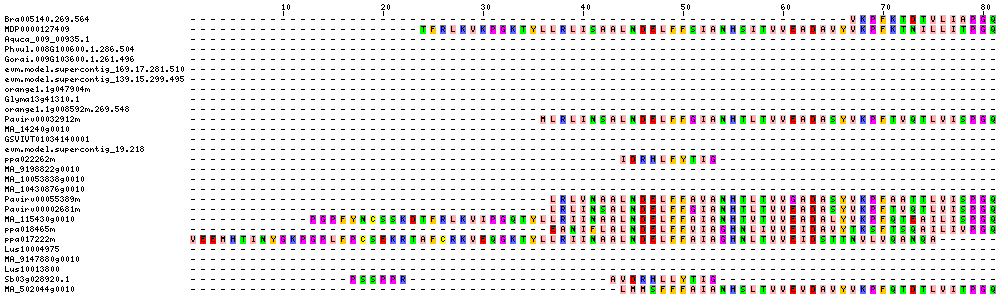
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |