

| Basic Information | |
|---|---|
| Species | Arabidopsis lyrata |
| Cazyme ID | 490996 |
| Family | AA4 |
| Protein Properties | Length: 560 Molecular Weight: 61535.7 Isoelectric Point: 7.3498 |
| Chromosome | Chromosome/Scaffold: 7 Start: 1845128 End: 1848857 |
| Description | FAD-linked oxidases family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA4 | 100 | 321 | 2.3e-22 |
| VSYFKEILGEKNVIEDKERLETANTDWMHKYKGSSKLMLLPKNTQEVSQILQYCDSRRLAVVPQGGNTGLVGGSVPVFDEVIINVGLMNKVLAFDEVSGV LVCEAGCILENLATFLDTKGFIMPLDLGAKGSCHIGGNVSTNAGGLRLIRYGSLHGTVLGLEAVTANGNVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIV TKVSILTQPKLSSVNLAFIACK | |||
| Full Sequence |
|---|
| Protein Sequence Length: 560 Download |
| MMMQKLRRSG ELIRFGCKSL FTSRPNKNLV SRSVSGFVNH YKSKGKLFEL SDGNYNTELH 60 HPCISRNLGM LLQQYKCFGS SAASKIQRNP LFSSLDSRDV SYFKEILGEK NVIEDKERLE 120 TANTDWMHKY KGSSKLMLLP KNTQEVSQIL QYCDSRRLAV VPQGGNTGLV GGSVPVFDEV 180 IINVGLMNKV LAFDEVSGVL VCEAGCILEN LATFLDTKGF IMPLDLGAKG SCHIGGNVST 240 NAGGLRLIRY GSLHGTVLGL EAVTANGNVL DMLGTLRKDN TGYDLKHLFI GSEGSLGIVT 300 KVSILTQPKL SSVNLAFIAC KDYLSCQKLL VEAKRNLGEI LSAFEFLDNN SMDLVLNHLD 360 GVRNPVSCSE NFYILIETTG SDETNDREKL EAFLLKSLEK GLVSDGVIAQ DINQASSFWR 420 IREGITEALQ KAGAVYKYDL SLPVEEIYNI VNDLRGKLGD LANVMGYGHL GDGNLHLNIS 480 AAEYNDKLLG LIEPYVYEWT SKHRGSISAE HGLGVMKANE IFYSKSPETV ALMASIKKLL 540 DPKGILNPYK VLPHSLFSH* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02805 | PLN02805 | 5.0e-38 | 100 | 553 | 469 | + D-lactate dehydrogenase [cytochrome] | ||
| PRK11230 | PRK11230 | 1.0e-39 | 100 | 553 | 478 | + glycolate oxidase subunit GlcD; Provisional | ||
| pfam02913 | FAD-oxidase_C | 7.0e-65 | 309 | 551 | 251 | + FAD linked oxidases, C-terminal domain. This domain has a ferredoxin-like fold. | ||
| TIGR00387 | glcD | 2.0e-77 | 138 | 550 | 430 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. | ||
| COG0277 | GlcD | 8.0e-109 | 102 | 554 | 464 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAB16815.1 | 0 | 36 | 559 | 4 | 524 | actin interacting protein [Arabidopsis thaliana] |
| RefSeq | NP_568003.2 | 0 | 1 | 559 | 1 | 559 | FAD linked oxidase family protein [Arabidopsis thaliana] |
| RefSeq | XP_002268002.1 | 0 | 78 | 558 | 69 | 550 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310828.1 | 0 | 73 | 559 | 46 | 529 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002521506.1 | 0 | 62 | 559 | 71 | 565 | d-lactate dehydrognease 2, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 0 | 94 | 552 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 0 | 94 | 552 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 0 | 94 | 552 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 0 | 94 | 552 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 0 | 94 | 552 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| GO838775 | 330 | 140 | 468 | 0 |
| GO839525 | 331 | 129 | 458 | 0 |
| CO071363 | 284 | 122 | 404 | 0 |
| FD574668 | 263 | 72 | 332 | 0 |
| CO071824 | 285 | 122 | 405 | 0 |
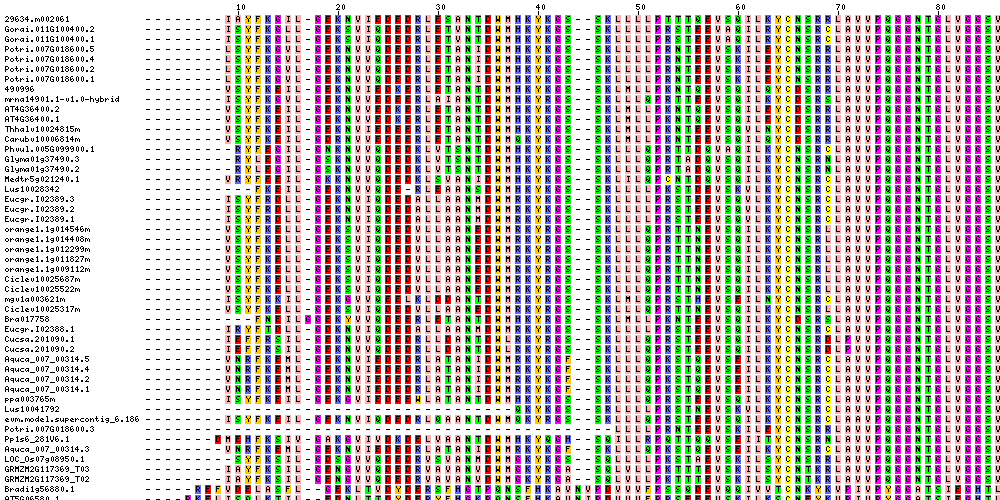
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |