

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_159128g0010 |
| Family | AA4 |
| Protein Properties | Length: 616 Molecular Weight: 65034.4 Isoelectric Point: 4.5869 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA4 | 11 | 240 | 3.6e-24 |
| ARRDEIVAALQKIVPGEGVISSAREMQPYESDALTAYRQPPMVVVLPDTTEQVSQVLKYCHDNGIKVVPRGSGTSLSGGSLPLADGVLLGLGKFKRVREI DFENRVAVVEPGVTNLAISQAVAHEGFYYAPDPSSQIACSIGGNIAENSGGVHSLKYGMTTNNVLGCEFVLITGEILRIGGKAPETDGYDVMGVITGSEG LLGVVTEVTVRILQKPETARALMVGFAEIE | |||
| Full Sequence |
|---|
| Protein Sequence Length: 616 Download |
| MMPEADQAVL ARRDEIVAAL QKIVPGEGVI SSAREMQPYE SDALTAYRQP PMVVVLPDTT 60 EQVSQVLKYC HDNGIKVVPR GSGTSLSGGS LPLADGVLLG LGKFKRVREI DFENRVAVVE 120 PGVTNLAISQ AVAHEGFYYA PDPSSQIACS IGGNIAENSG GVHSLKYGMT TNNVLGCEFV 180 LITGEILRIG GKAPETDGYD VMGVITGSEG LLGVVTEVTV RILQKPETAR ALMVGFAEIE 240 AAGQCVANII GAGIIPGGME MMDKNAIVAA EAFVHAGYPL DVEALLIIEL DGPKAEVDEL 300 IERVEKIARG CGSTTLQISN SEQERLLFWS GRKAAFPAVG RLSPDYLCMD GTIPRGKLPE 360 ALAGIRDLSE KYGLRVANVF HAGDGNLHPL ILYDANIAEE MDKAEAFGAD ILRLCVKLGG 420 VLTGEHGVGV EKRDLMPEMF SDIDLAQQQR LKCAFDTQGL LNPGKVFPTL HREPLAGSAS 480 HANETGQGVD TLKVRDAGDV EEAVRDALAA EQPLEIVGHG SKRAIGLPMA TNAVLDLSAL 540 NAVTSYEPNE LIVTVQAGAP MADLLSLIDS KNQQFAFDPM DTSVLLGTPR GAGTVGGMIA 600 AGLAGPRRIK AGGVRD |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0277 | GlcD | 4.0e-14 | 492 | 616 | 125 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| COG0277 | GlcD | 1.0e-111 | 19 | 470 | 462 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| TIGR00387 | glcD | 9.0e-165 | 54 | 465 | 413 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. | ||
| PRK11230 | PRK11230 | 0 | 13 | 472 | 460 | + glycolate oxidase subunit GlcD; Provisional | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_946481.1 | 0 | 1 | 472 | 4 | 475 | glycolate oxidase subunit GlcD [Rhodopseudomonas palustris CGA009] |
| RefSeq | YP_002290252.1 | 0 | 2 | 472 | 5 | 475 | glycolate oxidase, subunit GlcD [Oligotropha carboxidovorans OM5] |
| RefSeq | YP_485446.1 | 0 | 1 | 472 | 4 | 475 | FAD linked oxidase-like [Rhodopseudomonas palustris HaA2] |
| RefSeq | YP_534577.1 | 0 | 2 | 472 | 5 | 475 | FAD linked oxidase-like [Rhodopseudomonas palustris BisB18] |
| RefSeq | YP_571257.1 | 0 | 1 | 472 | 4 | 475 | D-lactate dehydrogenase (cytochrome) [Rhodopseudomonas palustris BisB5] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 9.80909e-45 | 15 | 466 | 16 | 475 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 9.80909e-45 | 15 | 466 | 16 | 475 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 9.80909e-45 | 15 | 466 | 16 | 475 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 9.80909e-45 | 15 | 466 | 16 | 475 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 9.80909e-45 | 15 | 466 | 16 | 475 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO388271 | 120 | 322 | 441 | 4.06377e-44 |
| GE179110 | 144 | 135 | 278 | 2.00386e-43 |
| BP853374 | 123 | 283 | 405 | 8.00001e-41 |
| CF306399 | 142 | 13 | 154 | 1e-33 |
| HO595273 | 270 | 207 | 468 | 1e-31 |
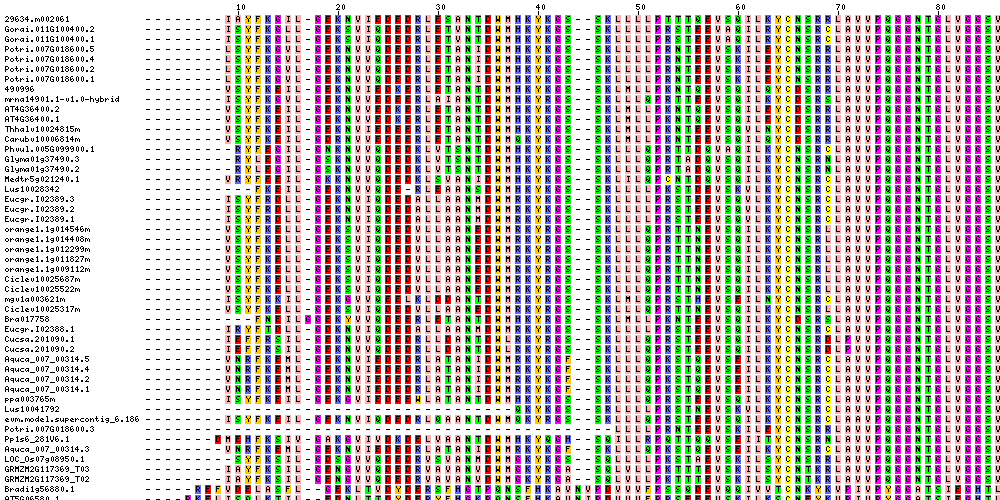
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |