

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s6_281V6.1 |
| Family | AA4 |
| Protein Properties | Length: 583 Molecular Weight: 63587.6 Isoelectric Point: 6.9798 |
| Chromosome | Chromosome/Scaffold: 6 Start: 2328573 End: 2334869 |
| Description | FAD-linked oxidases family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA4 | 106 | 288 | 9.1e-24 |
| DMEHFKSIVGAKGVIVDKDELVAANTDWMHKYQGHSQILLRPQTTQQVSEIITYCNSRNLAVVPQGGNTGLVGGSVPVFDEVIVNLGAMNKIIEFDEVSG ILVCEAGCILENLDNFIGGKGFTMPLDLGAKGSCQIGGNVSTNAGGLRLLRYGSLHGNVLGLEVVLADGEVVNMLGTLVKDNT | |||
| Full Sequence |
|---|
| Protein Sequence Length: 583 Download |
| MHPARRAADG LVRARLLGRW SWPLSFDCSI NRAAQRDALV SHRRLCTNRA RCLNKEYASV 60 NEGFFWTPEA EIGSLLKRHV KHQIRRFATS TKVNRNSRFD SLKTNDMEHF KSIVGAKGVI 120 VDKDELVAAN TDWMHKYQGH SQILLRPQTT QQVSEIITYC NSRNLAVVPQ GGNTGLVGGS 180 VPVFDEVIVN LGAMNKIIEF DEVSGILVCE AGCILENLDN FIGGKGFTMP LDLGAKGSCQ 240 IGGNVSTNAG GLRLLRYGSL HGNVLGLEVV LADGEVVNML GTLVKDNTGY DMKQLFIGSE 300 GTLGVVTKVS LLVPAKLGSV NTAFFACEDY TSCQNVLKEA KRQLGEILSA FEFIDRPALD 360 MALTHLPGTR DPLPQSGKNF YLLIETTGSN ETHDKEKLDA FLESTMEKGF VADGVVAQDS 420 TQAASFWQIR EGISEALGKA GAVYKYDFSL SPEHFYKIVE DLRDRLGSDA VVVGYGHLGD 480 GNVHLNVSAR EYKDELLEKV EPYVFEWVAK HKGSVSAEHG LGLMKPHALH YSKSAEAIAL 540 MERFKALLDP KGILNPYKVL PGTSKTVSTP ASALNAESAV VV* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02805 | PLN02805 | 7.0e-35 | 135 | 561 | 448 | + D-lactate dehydrogenase [cytochrome] | ||
| pfam01565 | FAD_binding_4 | 7.0e-38 | 142 | 278 | 137 | + FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidises the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. | ||
| pfam02913 | FAD-oxidase_C | 2.0e-59 | 319 | 559 | 249 | + FAD linked oxidases, C-terminal domain. This domain has a ferredoxin-like fold. | ||
| TIGR00387 | glcD | 4.0e-78 | 147 | 558 | 426 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. | ||
| COG0277 | GlcD | 2.0e-115 | 110 | 561 | 463 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| RefSeq | NP_568003.2 | 0 | 82 | 561 | 73 | 553 | FAD linked oxidase family protein [Arabidopsis thaliana] |
| RefSeq | XP_001752906.1 | 0 | 93 | 582 | 1 | 490 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_002268002.1 | 0 | 88 | 563 | 72 | 547 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310828.1 | 0 | 40 | 561 | 6 | 523 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002521506.1 | 0 | 75 | 561 | 72 | 559 | d-lactate dehydrognease 2, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 0 | 101 | 560 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 0 | 101 | 560 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 0 | 101 | 560 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 0 | 101 | 560 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 0 | 101 | 560 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| FC425720 | 243 | 292 | 534 | 0 |
| HO419275 | 406 | 160 | 564 | 0 |
| GO838775 | 330 | 147 | 476 | 0 |
| GO839525 | 331 | 136 | 466 | 0 |
| HO419275 | 34 | 130 | 163 | 0.043 |
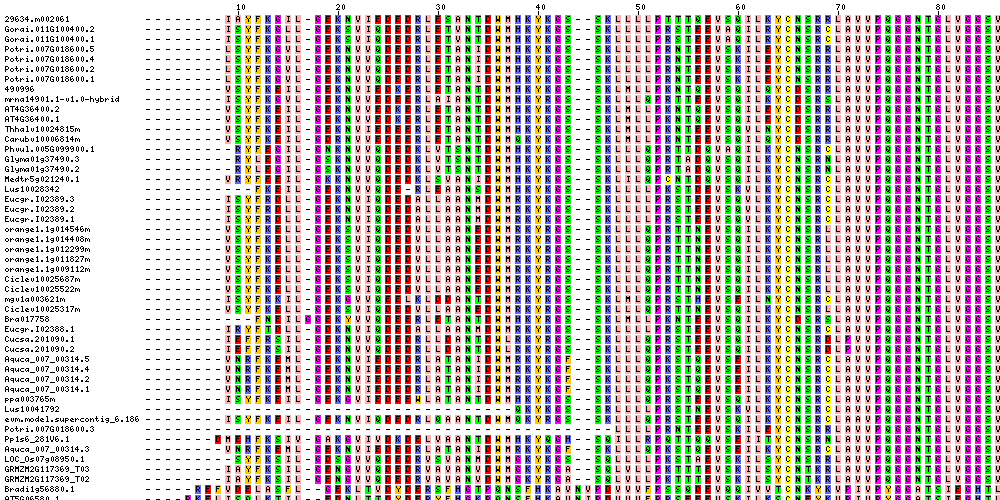
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |