

| Basic Information | |
|---|---|
| Species | Thellungiella halophila |
| Cazyme ID | Thhalv10024815m |
| Family | AA4 |
| Protein Properties | Length: 559 Molecular Weight: 61438.2 Isoelectric Point: 6.3031 |
| Chromosome | Chromosome/Scaffold: 1 Start: 1543706 End: 1547504 |
| Description | FAD-linked oxidases family protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA4 | 99 | 313 | 5.6e-23 |
| VSYFKEILGEKNVIEDEERLETANTDWMHKYKGSSKLMLLPKNTEEVSQVLNYCDSRRLAVVPQGGNTGLVGGSVPVFDEVIINVGLMNKVLAFDEVSGV LVCEAGCILENLANFLDTKGFVMPLDLGAKGSCHIGGNVSTNAGGLRLIRYGSLHGTVLGLEAVTANGNVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIV TKVSILTQPKLASVN | |||
| Full Sequence |
|---|
| Protein Sequence Length: 559 Download |
| MMMQKWRRSG DFIRFGRKSL TNLRSKEDSV SRSVSGFVNC CESKGKLFEP NAGNHNTELD 60 HRCPGRNLGI MQQYKCFGSS SASQIQRNPL FSSLDSNDVS YFKEILGEKN VIEDEERLET 120 ANTDWMHKYK GSSKLMLLPK NTEEVSQVLN YCDSRRLAVV PQGGNTGLVG GSVPVFDEVI 180 INVGLMNKVL AFDEVSGVLV CEAGCILENL ANFLDTKGFV MPLDLGAKGS CHIGGNVSTN 240 AGGLRLIRYG SLHGTVLGLE AVTANGNVLD MLGTLRKDNT GYDLKHLFIG SEGSLGIVTK 300 VSILTQPKLA SVNLAFIACK DYLSCQKILV EAKKNLGEIL SAFEFLDNNS MDLVLNHLDG 360 VRNPISSSEN FYILIETTGS DETSDREKLE AFLLKSLEKR LVSDGVIAQD INQASSFWRI 420 REGITEALQK AGAFYKYDLS LPVEEIYNIV NDLRGRLGDL ANVMGYGHLG DGNLHLNIST 480 VEYNDKILGL IEPYVYEWTS KHRGSISAEH GLGVMKANEI LYSKSPETVA LMASIKKLLD 540 PKGILNPYKV LPQSFFSP* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02805 | PLN02805 | 1.0e-37 | 99 | 552 | 469 | + D-lactate dehydrogenase [cytochrome] | ||
| PRK11230 | PRK11230 | 3.0e-41 | 99 | 552 | 489 | + glycolate oxidase subunit GlcD; Provisional | ||
| pfam02913 | FAD-oxidase_C | 3.0e-63 | 308 | 550 | 251 | + FAD linked oxidases, C-terminal domain. This domain has a ferredoxin-like fold. | ||
| TIGR00387 | glcD | 3.0e-77 | 137 | 549 | 430 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. | ||
| COG0277 | GlcD | 3.0e-111 | 101 | 552 | 463 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAB16815.1 | 0 | 36 | 557 | 4 | 523 | actin interacting protein [Arabidopsis thaliana] |
| RefSeq | NP_568003.2 | 0 | 1 | 557 | 1 | 558 | FAD linked oxidase family protein [Arabidopsis thaliana] |
| RefSeq | XP_002268002.1 | 0 | 56 | 557 | 54 | 550 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310828.1 | 0 | 65 | 557 | 42 | 528 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002521506.1 | 0 | 66 | 557 | 75 | 564 | d-lactate dehydrognease 2, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 0 | 93 | 551 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 0 | 93 | 551 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 0 | 93 | 551 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 0 | 93 | 551 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 0 | 93 | 551 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| GO838775 | 330 | 139 | 467 | 0 |
| GO839525 | 343 | 128 | 464 | 0 |
| FD574668 | 266 | 68 | 331 | 0 |
| CO071363 | 284 | 121 | 403 | 0 |
| HO419275 | 401 | 156 | 554 | 0 |
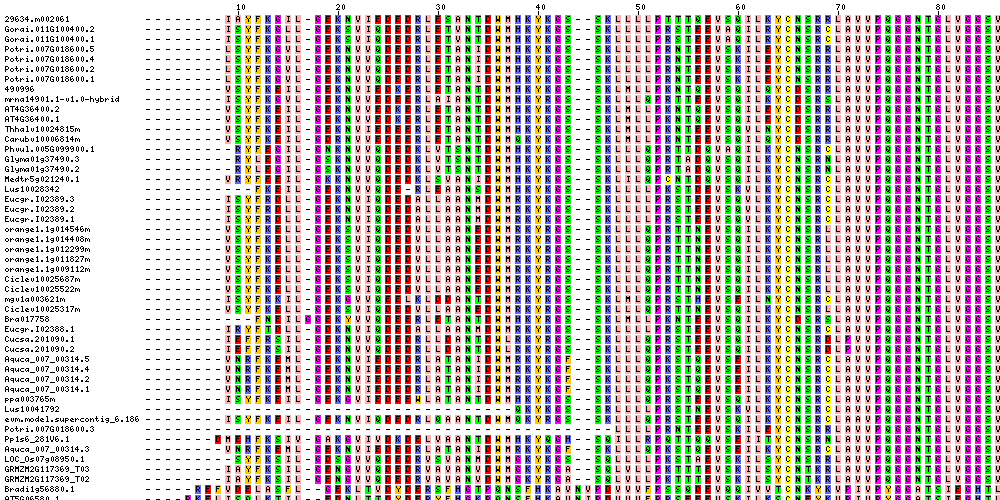
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |