

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s44_301V6.1 |
| Family | AA4 |
| Protein Properties | Length: 541 Molecular Weight: 59012.4 Isoelectric Point: 6.9076 |
| Chromosome | Chromosome/Scaffold: 44 Start: 1887221 End: 1894063 |
| Description | FAD-linked oxidases family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA4 | 66 | 280 | 1.5e-21 |
| SRLLAELKAVCDEDRVVVGVEERTFHAKPWHSHFAVKAIPDIVVYPKSQEEVVAVVNACAKYKVPMIPFAGGTSMEGQSMTPHKGVSINMKLMKRVKELH LEDMDVVVESGIGWIELNEFLKPHGLFFPLDPGPGASIGGMCATRCSGSLAVRYGTMRDNVISLKCILPNGDVVKTAARARKSAAGYDLTRLLIGSEGTL GLITEVTLRLQKIPE | |||
| Full Sequence |
|---|
| Protein Sequence Length: 541 Download |
| MGSGVTGWRK HARAIGGVVA FTGLMSLLKL HEWVASRIED LEQVEATERP VSWNDWSRNL 60 IVDIISRLLA ELKAVCDEDR VVVGVEERTF HAKPWHSHFA VKAIPDIVVY PKSQEEVVAV 120 VNACAKYKVP MIPFAGGTSM EGQSMTPHKG VSINMKLMKR VKELHLEDMD VVVESGIGWI 180 ELNEFLKPHG LFFPLDPGPG ASIGGMCATR CSGSLAVRYG TMRDNVISLK CILPNGDVVK 240 TAARARKSAA GYDLTRLLIG SEGTLGLITE VTLRLQKIPE ASVKIALKPP QTGAVQVAMC 300 NFKSIGDATA VAIATMHSGI QVSRVELLDE MMMNAINIAN GKNYPTEPTL MFEFVGTEAY 360 ALEQTSRVEK IVQAHNGSDF VYADSEMEKE ELWKIRKGAF WSSFSLRPGA EGMTTDVCVP 420 LSRLAECIST SKERLLASSL PHVLVAHAGD GNFHVCIFFD KNNEEEVKEA LEIAPPPPPS 480 FVSKLSRTCT GEHGVGVGKM KYLEKEHGSA AMTMMGSIKR AIDPSNLMNP GKLIPEKFCY 540 * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK11230 | PRK11230 | 2.0e-47 | 100 | 535 | 447 | + glycolate oxidase subunit GlcD; Provisional | ||
| pfam02913 | FAD-oxidase_C | 7.0e-58 | 294 | 533 | 249 | + FAD linked oxidases, C-terminal domain. This domain has a ferredoxin-like fold. | ||
| TIGR00387 | glcD | 2.0e-76 | 108 | 532 | 436 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. | ||
| COG0277 | GlcD | 5.0e-103 | 101 | 537 | 454 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| PLN02805 | PLN02805 | 0 | 90 | 540 | 451 | + D-lactate dehydrogenase [cytochrome] | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABK24997.1 | 0 | 45 | 540 | 95 | 580 | unknown [Picea sitchensis] |
| RefSeq | XP_001755583.1 | 0 | 112 | 540 | 6 | 421 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001760719.1 | 0 | 131 | 540 | 1 | 397 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_002322755.1 | 0 | 66 | 540 | 20 | 480 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002524111.1 | 0 | 67 | 540 | 97 | 555 | d-lactate dehydrogenase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 5e-38 | 107 | 534 | 54 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 5e-38 | 107 | 534 | 54 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 5e-38 | 107 | 534 | 54 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 5e-38 | 107 | 534 | 54 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 5e-38 | 107 | 534 | 54 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| photorespiration | RXN-969 | EC-1.1.3 | glycolate oxidase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| FC436506 | 245 | 297 | 541 | 0 |
| FC436505 | 227 | 315 | 541 | 0 |
| EX265038 | 337 | 120 | 453 | 0 |
| EX440808 | 335 | 90 | 424 | 0 |
| DY305587 | 311 | 159 | 469 | 0 |
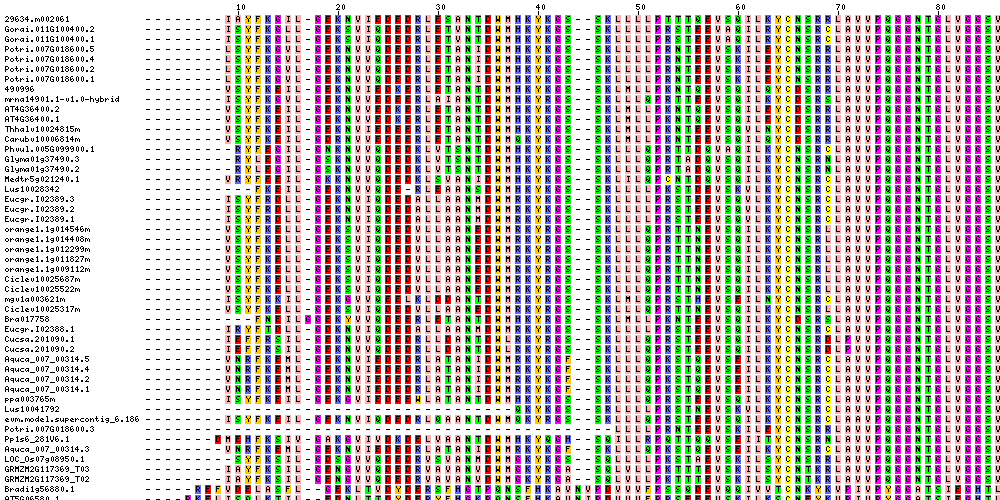
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |