

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.011G100400.2 |
| Family | AA4 |
| Protein Properties | Length: 542 Molecular Weight: 59673.4 Isoelectric Point: 6.5833 |
| Chromosome | Chromosome/Scaffold: 11 Start: 11172087 End: 11180531 |
| Description | FAD-linked oxidases family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA4 | 82 | 288 | 8.4e-23 |
| ISYFKGLLGEKSVIQDEDRLETVNTDWMHKYKGSSKLLLLPRSTEEVAQILRYCNSRCLAVVPQGGNTGLVGGSVPVFDEVIVNISSMNNIISFDKVSGI LVCEAGCILENLISFLDNQGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGNVLGLEAVLANGDVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIV TKVSILT | |||
| Full Sequence |
|---|
| Protein Sequence Length: 542 Download |
| MEKWRATSNL RRSLKSILNR QLSSVSEFRY LNEKRSCQSS FNLIRDCKSL GQVNAIQHRC 60 FSSASTLVQR NPSFSTLNSD DISYFKGLLG EKSVIQDEDR LETVNTDWMH KYKGSSKLLL 120 LPRSTEEVAQ ILRYCNSRCL AVVPQGGNTG LVGGSVPVFD EVIVNISSMN NIISFDKVSG 180 ILVCEAGCIL ENLISFLDNQ GFIMPLDLGA KGSCQIGGNV STNAGGLRLV RYGSLHGNVL 240 GLEAVLANGD VLDMLGTLRK DNTGYDLKHL FIGSEGSLGI VTKVSILTPP KLSSVNIAFL 300 ACNDYSSCQK LLMEAKRKLG EILSAFEFLD TEAMNLVLHQ LDGVRNPLPA SMHNFYILIE 360 TTGSDESYNR EKLEAFLLSS MEGGLISDGV LAQDINQASS FWRIREGVPE ALMKAGAVYK 420 YDLSLPVEKM YDLVDDMRIR LGDLAKVVGY GHLGDGNLHL NVSAPEYDDK ILEQIEPYVY 480 EWTSKHRGSI SAEHGLGLMK ANKIYYSKSA ETVQTMASIK KLLDPNGILN PYKVLPHLLN 540 S* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK11230 | PRK11230 | 1.0e-34 | 94 | 538 | 481 | + glycolate oxidase subunit GlcD; Provisional | ||
| PLN02805 | PLN02805 | 1.0e-40 | 110 | 536 | 446 | + D-lactate dehydrogenase [cytochrome] | ||
| pfam02913 | FAD-oxidase_C | 5.0e-59 | 291 | 534 | 252 | + FAD linked oxidases, C-terminal domain. This domain has a ferredoxin-like fold. | ||
| TIGR00387 | glcD | 8.0e-70 | 120 | 533 | 430 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. | ||
| COG0277 | GlcD | 6.0e-104 | 84 | 538 | 466 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAB16815.1 | 0 | 57 | 541 | 42 | 523 | actin interacting protein [Arabidopsis thaliana] |
| RefSeq | NP_568003.2 | 0 | 57 | 541 | 74 | 558 | FAD linked oxidase family protein [Arabidopsis thaliana] |
| RefSeq | XP_002268002.1 | 0 | 51 | 541 | 60 | 550 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310828.1 | 0 | 1 | 541 | 1 | 528 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002521506.1 | 0 | 20 | 541 | 43 | 564 | d-lactate dehydrognease 2, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 0 | 76 | 535 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 0 | 76 | 535 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 0 | 76 | 535 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 0 | 76 | 535 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 0 | 76 | 535 | 12 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| ES812965 | 298 | 2 | 299 | 0 |
| CO071363 | 284 | 104 | 387 | 0 |
| CO071824 | 284 | 105 | 388 | 0 |
| GO838775 | 330 | 122 | 451 | 0 |
| ES812965 | 52 | 299 | 350 | 0.007 |
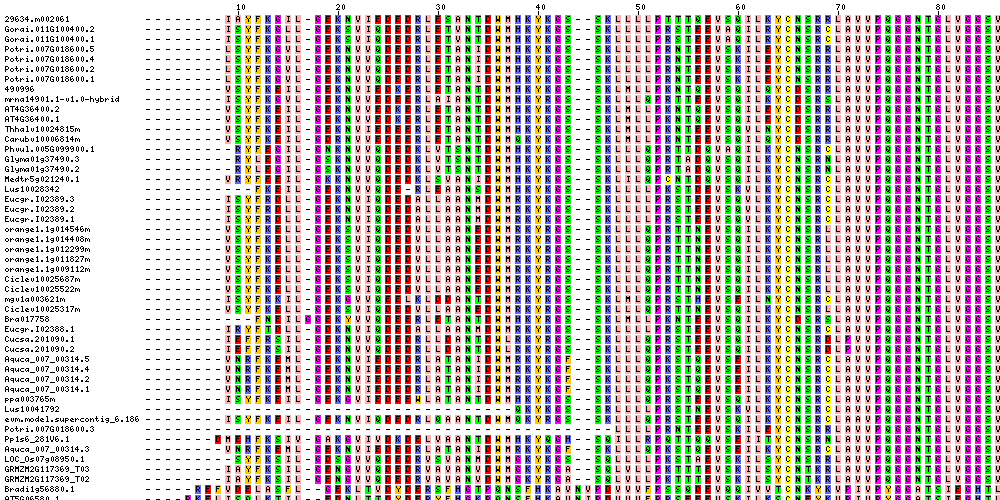
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |