

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_10433252g0020 |
| Family | PL1 |
| Protein Properties | Length: 309 Molecular Weight: 34517.3 Isoelectric Point: 6.8169 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL1 | 157 | 295 | 1.6e-34 |
| VIQHEPLWIIFESDMVIRLKEELLMNSFKTIDGRGADVHIAHGACITIQFVTNIIIHGVSIHDCIQTGNAMVRDSPDHYGFRTLADGDGISIFGGRFIWI DHCSFSSCHHGLIDAIMGSTAITISNNYFTDHNMVSCIT | |||
| Full Sequence |
|---|
| Protein Sequence Length: 309 Download |
| MTHQSQLHYL FLFVGIVCLQ IFAPKCSAAK TQGMLDSIMG FLRGRPTIQA AQQETRHHKH 60 NSSFPVESPE EVVKMVQKNI NDSRRQLTYL SCGTGNPIDD CWRCDPNWQM NRQRLADCAI 120 GFGRNAIGGK NGQYYVVTDS SDNDAVNPTP GTLRHAVIQH EPLWIIFESD MVIRLKEELL 180 MNSFKTIDGR GADVHIAHGA CITIQFVTNI IIHGVSIHDC IQTGNAMVRD SPDHYGFRTL 240 ADGDGISIFG GRFIWIDHCS FSSCHHGLID AIMGSTAITI SNNYFTDHNM VSCITMTLDS 300 FYKSSVLVY |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG3866 | PelB | 8.0e-11 | 128 | 302 | 186 | + Pectate lyase [Carbohydrate transport and metabolism] | ||
| pfam00544 | Pec_lyase_C | 2.0e-25 | 176 | 291 | 138 | + Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. | ||
| smart00656 | Amb_all | 1.0e-33 | 170 | 292 | 134 | + Amb_all domain. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABK25017.1 | 0 | 9 | 291 | 4 | 283 | unknown [Picea sitchensis] |
| GenBank | ABR18471.1 | 0 | 1 | 307 | 1 | 303 | unknown [Picea sitchensis] |
| EMBL | CAN76500.1 | 0 | 62 | 291 | 21 | 250 | hypothetical protein [Vitis vinifera] |
| EMBL | CBI14899.1 | 0 | 62 | 291 | 21 | 250 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002275781.1 | 0 | 62 | 291 | 76 | 305 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1pxz_B | 0 | 96 | 291 | 2 | 198 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1pxz_A | 0 | 96 | 291 | 2 | 198 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 3zsc_A | 0.00005 | 165 | 294 | 50 | 167 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| PDB | 3krg_A | 0.0003 | 244 | 292 | 184 | 249 | A Chain A, Structural Insights Into Substrate Specificity And The Anti Beta-Elimination Mechanism Of Pectate Lyase |
| PDB | 2o1d_A | 0.0003 | 244 | 292 | 184 | 249 | A Chain A, Structural Insights Into Substrate Specificity And The Anti Beta-Elimination Mechanism Of Pectate Lyase |
| Transmembrane Domains | ||||
|---|---|---|---|---|
 | ||||
| Start | End | |||
| 7 | 24 | |||
| Signal Peptide | ||||
|---|---|---|---|---|
 | ||||
| Cleavage Site | ||||
| 0 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR479712 | 272 | 1 | 272 | 0 |
| EX393983 | 287 | 21 | 307 | 0 |
| DR485347 | 206 | 1 | 206 | 0 |
| GW756557 | 232 | 1 | 232 | 0 |
| DR523820 | 231 | 1 | 231 | 0 |
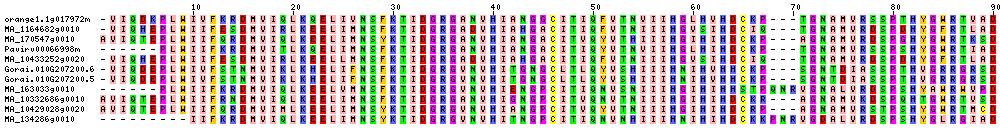
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |