

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_163033g0010 |
| Family | PL1 |
| Protein Properties | Length: 244 Molecular Weight: 27723.5 Isoelectric Point: 6.6651 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL1 | 99 | 232 | 2.2e-34 |
| PLWIIFKRDMVIQLKEELVMNSFKTIDGRGVNVHIENGPCITIQNVSNIIIHGIHIHHSTPQNRVGNALVRDSPSHYAWRWVPDGNGISIIGSTQIWIDH CSLSKCRDGLIDAVSGSTLITISNNYFTDHDDVS | |||
| Full Sequence |
|---|
| Protein Sequence Length: 244 Download |
| MIYASFLYDV IYHCRNITNI ARRELAFLSQ GTGNPVDDCW CLDPNWRRNR KLLADCGVGF 60 GKEAIGGKEG RIYVVRDPRD DPMNPWGDTL RYAVIQEEPL WIIFKRDMVI QLKEELVMNS 120 FKTIDGRGVN VHIENGPCIT IQNVSNIIIH GIHIHHSTPQ NRVGNALVRD SPSHYAWRWV 180 PDGNGISIIG STQIWIDHCS LSKCRDGLID AVSGSTLITI SNNYFTDHDD VSQSIPSYVF 240 LHQF |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG3866 | PelB | 3.0e-11 | 162 | 232 | 80 | + Pectate lyase [Carbohydrate transport and metabolism] | ||
| pfam00544 | Pec_lyase_C | 4.0e-25 | 113 | 231 | 142 | + Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. | ||
| smart00656 | Amb_all | 3.0e-27 | 107 | 232 | 137 | + Amb_all domain. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK66161.1 | 0 | 22 | 231 | 8 | 215 | pectate lyase [Fragaria x ananassa] |
| GenBank | EEE60442.1 | 0 | 15 | 231 | 78 | 290 | hypothetical protein OsJ_13660 [Oryza sativa Japonica Group] |
| RefSeq | NP_001141589.1 | 0 | 9 | 231 | 95 | 313 | hypothetical protein LOC100273705 [Zea mays] |
| RefSeq | NP_001147553.1 | 0 | 9 | 231 | 109 | 327 | pectate lyase 8 [Zea mays] |
| RefSeq | XP_002446075.1 | 0 | 9 | 231 | 81 | 299 | hypothetical protein SORBIDRAFT_06g001410 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1pxz_B | 0 | 34 | 231 | 2 | 198 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1pxz_A | 0 | 34 | 231 | 2 | 198 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 3zsc_A | 0.0000002 | 65 | 232 | 22 | 165 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| PDB | 1vbl_A | 0.00003 | 116 | 232 | 128 | 255 | A Chain A, Structure Of The Thermostable Pectate Lyase Pl 47 |
| PDB | 3krg_A | 0.0001 | 179 | 232 | 179 | 249 | A Chain A, Structural Insights Into Substrate Specificity And The Anti Beta-Elimination Mechanism Of Pectate Lyase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR683957 | 205 | 19 | 223 | 0 |
| DR686230 | 193 | 19 | 211 | 0 |
| DT633426 | 191 | 19 | 209 | 0 |
| FL748690 | 224 | 9 | 231 | 0 |
| DV984437 | 214 | 19 | 231 | 0 |
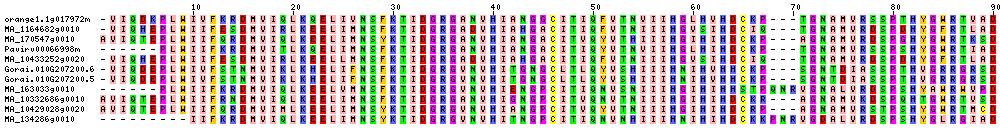
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |