

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_134286g0010 |
| Family | PL1 |
| Protein Properties | Length: 276 Molecular Weight: 30918.2 Isoelectric Point: 7.7167 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL1 | 126 | 255 | 1.6e-35 |
| IIFKRDMVIQLKEELIMNSYKTIDGRGVNVHITNGPCITIQNVNHIIIHNIHIHDCKKPNRVGDALVRDSPSHYGLRGIADQDGISILASTHIWIDHCSL SKCDDGLIDAVSASTAITISNNYFTDHDKV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 276 Download |
| MLLVGILYIC SQAPQCFATK AFGGEAAPAN NSFISVAEKS NINSTRRELA YFSYATGNPI 60 DDCWRWDSNW RRNRKRLADC GVGFGKDAIG GKEGRIYVVR DPRDDPVNPW RGTLRYGVIQ 120 DEPLWIIFKR DMVIQLKEEL IMNSYKTIDG RGVNVHITNG PCITIQNVNH IIIHNIHIHD 180 CKKPNRVGDA LVRDSPSHYG LRGIADQDGI SILASTHIWI DHCSLSKCDD GLIDAVSAST 240 AITISNNYFT DHDKVPHSIP SYVFLPSLCI NSRFQC |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG3866 | PelB | 5.0e-10 | 208 | 255 | 57 | + Pectate lyase [Carbohydrate transport and metabolism] | ||
| pfam00544 | Pec_lyase_C | 7.0e-24 | 137 | 255 | 142 | + Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. | ||
| smart00656 | Amb_all | 6.0e-25 | 131 | 255 | 136 | + Amb_all domain. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAB10560.1 | 0 | 35 | 255 | 37 | 255 | pectate lyase [Arabidopsis thaliana] |
| RefSeq | NP_001141589.1 | 0 | 44 | 255 | 104 | 313 | hypothetical protein LOC100273705 [Zea mays] |
| RefSeq | NP_568967.1 | 0 | 36 | 255 | 60 | 277 | pectate lyase family protein [Arabidopsis thaliana] |
| RefSeq | XP_002297822.1 | 0 | 25 | 255 | 66 | 297 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002446075.1 | 0 | 27 | 255 | 77 | 299 | hypothetical protein SORBIDRAFT_06g001410 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1pxz_B | 0 | 58 | 255 | 2 | 198 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1pxz_A | 0 | 58 | 255 | 2 | 198 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 3zsc_A | 0.000000003 | 77 | 255 | 2 | 164 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| PDB | 1vbl_A | 0.000003 | 140 | 255 | 128 | 254 | A Chain A, Structure Of The Thermostable Pectate Lyase Pl 47 |
| PDB | 2bsp_A | 0.00003 | 147 | 254 | 151 | 268 | A Chain A, Bacillus Subtilis Pectate Lyase R279k Mutant |
| Signal Peptide | ||||
|---|---|---|---|---|
 | ||||
| Cleavage Site | ||||
| 0 | ||||
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR683957 | 247 | 1 | 247 | 0 |
| DR686230 | 235 | 1 | 235 | 0 |
| DT633426 | 233 | 1 | 233 | 0 |
| DT631978 | 228 | 1 | 228 | 0 |
| DV984437 | 256 | 1 | 255 | 0 |
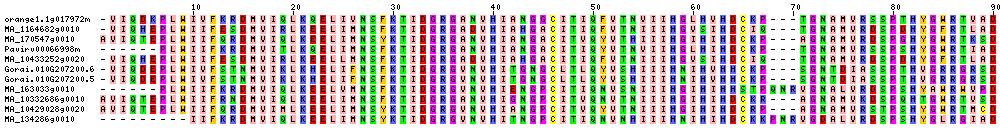
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |