

| Basic Information | |
|---|---|
| Species | Linum usitatissimum |
| Cazyme ID | Lus10002096 |
| Family | GT48 |
| Protein Properties | Length: 1168 Molecular Weight: 130074 Isoelectric Point: 9.767 |
| Chromosome | Chromosome/Scaffold: 575 Start: 62053 End: 70031 |
| Description | callose synthase 5 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT48 | 106 | 504 | 0 |
| IRFHYGHPDVFDRIFHITRGGVSKASRGINLSEDIFAGFNSTLRRGNITHHEYIQVGKGRDVGLNQISLFEAKVACGNGEQTLSRDIYRLGHRFDFFRML SCYYTTIGFYISSMVVVLTVYAFLYGRIYLALSGLEGSIIKFAKHRGDTALKSAMASQSVVQLGILTALPMVMEMGLERGFRTALGDIIIMQLQLAAVFF TFSLGTRVHYFGRTVLHGGAKYRAIGRGFVVRHEKFAENYRMYSRSHFVKALELMLLLICYQIYGKAVNDQVAYMLLTASMWFLVSSWLFAPFLFNPSGF EWQKIVDDWDDWTKWINSRGGIGVPANKSWESWWEEEQEHLQHTGISGQICEVILALRFFIYQYGIVYQLKVTKATSAGREHSISVYGLSWLVIVALML | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1168 Download |
| MEERDRGKSQ KVYYSVLVKA VDNLDQEIYR IKLPGQAKLG EGKPENQNHA IVFTRGEAMQ 60 AIDMNQDNYL EEAFKMRNLL EEFHEDHGVR PPTILGVREH IFTGGIRFHY GHPDVFDRIF 120 HITRGGVSKA SRGINLSEDI FAGFNSTLRR GNITHHEYIQ VGKGRDVGLN QISLFEAKVA 180 CGNGEQTLSR DIYRLGHRFD FFRMLSCYYT TIGFYISSMV VVLTVYAFLY GRIYLALSGL 240 EGSIIKFAKH RGDTALKSAM ASQSVVQLGI LTALPMVMEM GLERGFRTAL GDIIIMQLQL 300 AAVFFTFSLG TRVHYFGRTV LHGGAKYRAI GRGFVVRHEK FAENYRMYSR SHFVKALELM 360 LLLICYQIYG KAVNDQVAYM LLTASMWFLV SSWLFAPFLF NPSGFEWQKI VDDWDDWTKW 420 INSRGGIGVP ANKSWESWWE EEQEHLQHTG ISGQICEVIL ALRFFIYQYG IVYQLKVTKA 480 TSAGREHSIS VYGLSWLVIV ALMLILKTVS TGRKKFSADF QLMFRLLKFI LFIGFVVTLI 540 ILFTTLHLTV GDVFQSLLAF LPTGWAILQI SQACKPIVKG MKMWGSVKAL GRGYEYMMGL 600 ILFLPIAVLA WFPFVSEFQN RLLFNQAFSR GLQIQRILAG GKKHKSPGNP GGRAEAKAYM 660 GGSRSPGPGS RSPCPGSRSP GPRSRSPVPG SRGASRPTSP LHPRLSGLNL DSPTGRGSED 720 GKSQCHPLPL PPTSPTSPSS SVTSTRSPGA TEGAAATLSR WKKGKLLGRG TFGQVYLGFN 780 SVNGQMCAIK EVRVVSDDQT SKECLKQLNQ EINLLSQLSH PNIVKYHGSE LSEETLSVYL 840 EFVSGGSIHK LLQEYGAFKE PVIQNYTRQI LSGLAYLHGR NTVHRDIKGA NILVDPNGEI 900 KLADFGMAKH ITACSSMLSF KGSPYWMAPE VVMNTNGYSL AVDIWSLGCT ILEMAAGKPP 960 WSQYEGVAAI FKIGNSKDMP EIPYFLSDEA KSFIKLCLQR DPSLRPTASQ LLDHPFLRDH 1020 AMRVPNSNPS SPYNFDGSRT PPALDLQPNR NGITSHEAEY PIKHVITNAR GTRTSRDDVR 1080 MITSLPVSPC SSPLRQNGPA HRSCFLSPPH LYPLMGQNGS YNNVSDFLLQ PVRQPPALSH 1140 SDFVLDSNSL MKQMTGPSSQ PIIRRNS* |
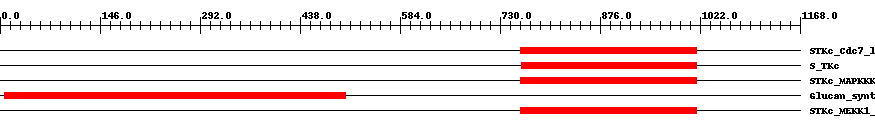
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| cd06627 | STKc_Cdc7_like | 3.0e-95 | 760 | 1017 | 259 | + Catalytic domain of Cell division control protein 7-like Protein Serine/Threonine Kinases. Serine/threonine kinases (STKs), (Cdc7)-like subfamily, catalytic (c) domain. STKs catalyze the transfer of the gamma-phosphoryl group from ATP to serine/threonine residues on protein substrates. The Cdc7-like subfamily is part of a larger superfamily that includes the catalytic domains of other protein STKs, protein tyrosine kinases, RIO kinases, aminoglycoside phosphotransferase, choline kinase, and phosphoinositide 3-kinase. Members of this subfamily include Schizosaccharomyces pombe Cdc7, Saccharomyces cerevisiae Cdc15, Arabidopsis thaliana mitogen-activated protein kinase (MAPK) kinase kinase (MAPKKK) epsilon, and related proteins. MAPKKKs phosphorylate and activate MAPK kinases (MAPKKs or MKKs or MAP2Ks), which in turn phosphorylate and activate MAPKs during signaling cascades that are important in mediating cellular responses to extracellular signals. Fission yeast Cdc7 is essential for cell division by playing a key role in the initiation of septum formation and cytokinesis. Budding yeast Cdc15 functions to coordinate mitotic exit with cytokinesis. Arabidopsis MAPKKK epsilon is required for pollen development in the plasma membrane. | ||
| smart00220 | S_TKc | 3.0e-95 | 761 | 1017 | 259 | + Serine/Threonine protein kinases, catalytic domain. Phosphotransferases. Serine or threonine-specific kinase subfamily. | ||
| cd06606 | STKc_MAPKKK | 7.0e-127 | 760 | 1017 | 264 | + Catalytic domain of the Protein Serine/Threonine Kinase, Mitogen-Activated Protein Kinase Kinase Kinase. Serine/threonine kinases (STKs), mitogen-activated protein kinase (MAPK) kinase kinase (MAPKKK) subfamily, catalytic (c) domain. STKs catalyze the transfer of the gamma-phosphoryl group from ATP to serine/threonine residues on protein substrates. The MAPKKK subfamily is part of a larger superfamily that includes the catalytic domains of other protein STKs, protein tyrosine kinases, RIO kinases, aminoglycoside phosphotransferase, choline kinase, and phosphoinositide 3-kinase. MAPKKKs (MKKKs or MAP3Ks) are also called MAP/ERK kinase kinases (MEKKs) in some cases. They phosphorylate and activate MAPK kinases (MAPKKs or MKKs or MAP2Ks), which in turn phosphorylate and activate MAPKs during signaling cascades that are important in mediating cellular responses to extracellular signals. This subfamily is composed of the Apoptosis Signal-regulating Kinases ASK1 (or MAPKKK5) and ASK2 (or MAPKKK6), MEKK1, MEKK2, MEKK3, MEKK4, as well as plant and fungal MAPKKKs. Also included in this subfamily are the cell division control proteins Schizosaccharomyces pombe Cdc7 and Saccharomyces cerevisiae Cdc15. | ||
| pfam02364 | Glucan_synthase | 1.0e-159 | 7 | 505 | 576 | + 1,3-beta-glucan synthase component. This family consists of various 1,3-beta-glucan synthase components including Gls1, Gls2 and Gls3 from yeast. 1,3-beta-glucan synthase EC:2.4.1.34 also known as callose synthase catalyzes the formation of a beta-1,3-glucan polymer that is a major component of the fungal cell wall. The reaction catalyzed is:- UDP-glucose + {(1,3)-beta-D-glucosyl}(N) <=> UDP + {(1,3)-beta-D-glucosyl}(N+1). | ||
| cd06632 | STKc_MEKK1_plant | 9.0e-160 | 760 | 1017 | 258 | + Catalytic domain of the Protein Serine/Threonine Kinase, Plant MAP/ERK kinase kinase 1. Serine/threonine kinases (STKs), plant MAP/ERK kinase kinase 1 (MEKK1)-like subfamily, catalytic (c) domain. STKs catalyze the transfer of the gamma-phosphoryl group from ATP to serine/threonine residues on protein substrates. The plant MEKK1 subfamily is part of a larger superfamily that includes the catalytic domains of other protein STKs, protein tyrosine kinases, RIO kinases, aminoglycoside phosphotransferase, choline kinase, and phosphoinositide 3-kinase. This subfamily is composed of plant mitogen-activated protein kinase (MAPK) kinase kinases (MAPKKKs or MKKKs or MAP3Ks) including Arabidopsis thaliana MEKK1 and MAPKKK3. MEKK1 is a MAPKKK that phosphorylates and activates MAPK kinases (MAPKKs or MKKs or MAP2Ks), which in turn phosphorylate and activate MAPKs during signaling cascades that are important in mediating cellular responses to extracellular signals. Arabidopsis thaliana MEKK1 activates MPK4, a MAPK that regulates systemic acquired resistance. MEKK1 also participates in the regulation of temperature-sensitive and tissue-specific cell death. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0000148 | 1,3-beta-D-glucan synthase complex |
| GO:0003843 | 1,3-beta-D-glucan synthase activity |
| GO:0004672 | protein kinase activity |
| GO:0005524 | ATP binding |
| GO:0006075 | (1->3)-beta-D-glucan biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK49452.2 | 0 | 1 | 645 | 1264 | 1931 | AF304372_1 putative beta-1,3-glucan synthase [Nicotiana alata] |
| GenBank | AAM15250.1 | 0 | 1 | 645 | 214 | 878 | putative 1,3-beta-D-glucan synthase [Arabidopsis thaliana] |
| GenBank | AAM15369.1 | 0 | 1 | 645 | 214 | 878 | putative 1,3-beta-D-glucan synthase [Arabidopsis thaliana] |
| RefSeq | XP_002328472.1 | 0 | 8 | 645 | 1242 | 1906 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002529727.1 | 0 | 1 | 645 | 1192 | 1864 | transferase, transferring glycosyl groups, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3vw6_B | 0 | 766 | 1018 | 15 | 269 | A Chain A, Crystal Structure Of Human Apoptosis Signal-Regulating Kinase 1 (Ask1) With Imidazopyridine Inhibitor |
| PDB | 3vw6_A | 0 | 766 | 1018 | 15 | 269 | A Chain A, Crystal Structure Of Human Apoptosis Signal-Regulating Kinase 1 (Ask1) With Imidazopyridine Inhibitor |
| PDB | 3a7j_A | 0 | 748 | 1024 | 11 | 281 | A Chain A, Crystal Structure Of Human Apoptosis Signal-Regulating Kinase 1 (Ask1) With Imidazopyridine Inhibitor |
| PDB | 3a7i_A | 0 | 748 | 1024 | 11 | 281 | A Chain A, Crystal Structure Of Human Apoptosis Signal-Regulating Kinase 1 (Ask1) With Imidazopyridine Inhibitor |
| PDB | 3a7h_B | 0 | 748 | 1024 | 11 | 281 | A Chain A, Crystal Structure Of Human Apoptosis Signal-Regulating Kinase 1 (Ask1) With Imidazopyridine Inhibitor |