

| Basic Information | |
|---|---|
| Species | Linum usitatissimum |
| Cazyme ID | Lus10002097 |
| Family | GT48 |
| Protein Properties | Length: 1199 Molecular Weight: 138270 Isoelectric Point: 9.3942 |
| Chromosome | Chromosome/Scaffold: 575 Start: 74958 End: 83588 |
| Description | callose synthase 5 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT48 | 995 | 1193 | 0 |
| VPANLEARRRIAFFTNSLFMDMPRAPRVRKMLSFSVMTPYYSEETVYSKSDLEMENEDDEWTNFMERINCKKESEVWENEENILQLRHWASLRGQTLCRT LRGMMYYRRALKLQAFLDMANESDMKLVSDFRYAEILAGYKAITNPSEEDKKSQRSLSAQLEAVADMKFTYVATCQIYGNQKRKGDRHATNILNLMVKY | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1199 Download |
| MINTEAYPHL IRRPSRSAAT TTFSTEVFDN SVVPSSLESI KPILRTANEI QNERPRVAYL 60 CRFYAFEKAH RLDPSSSGRG VRQFKTALLQ RVERENASSI ASRVKKTDAR EIQSFYYQYY 120 KQYVEALEQG EQADRAQLGK VYETAGVLYE VLCAVNKAEK VEEVAPEIIA AARDIQEKKE 180 IYAPYNILPL DSAGASQSIM QLEEVKAAVA ALWNTRGLSW PSSFEQHRQK AGDLDLLDWL 240 RAMFGFQRDS VRNQREHLIL LLANNHIRLN PKPEPLNKAR AVPDLDERAV DAVMQKIFKN 300 YKNWCKFLGR KHSLRLPQSQ PEIQQRKILY MGLYLLIWGE AANVRFMPEC LCYIFHNMAY 360 ELHGLLAGNV SIVTGENIKP SYGGDDEAFL RKVIAPVYRV IAKEASKSQN GMASSTEWCN 420 YDDLNEYFWS SDCFSLGWPM RDDGEFFKST RDAAKGKKTS QENSGSTGKS YFVETRAFWH 480 IFRSFDRLWT FYVLAFQLMI IYAWSGVAIQ NILRRDVLYY LSSIFITAAY LRLLQSILDL 540 VLNFPGYHRW KFTDVLRNIL KIIVSFAWAI ILPLCYSGNF KQVGGFTKGM NFLRTVKSIP 600 PIYLFSVVIY MIPNILAACL FIFPMLRRWI ENSDWLIVRL LLWWSQPRIY VGRGMHESQF 660 SLIKYTIFWV LLLASKFTFS FFIQIKPLVK PTKDIMNIRH VEYTWHELFP YARHNYGAVL 720 SLWAPTILIR TLGMLRSRFQ SLPGAFNAYL VPSDKMRKKG FSLSKRFADV TANRRSEAAK 780 FAQLEMDLLL VPYTSDPSIK LIQWPPFLLA SKIPIALNMA AQFRSKDSDL WKRICADEYM 840 KCAVIECYES FKQVLNILIV GENEKRIIGI IIKEIESHVS KGTLLANFRM ASLPPLCEKV 900 VVLVGILKDG HPSERDNVVL LLQDMLELVT RDLMVNENRE LVDIGHSGKD SGRQIFAGTD 960 PRPAITFPPP VSAQWDEQIR RLHLLLTVKE SAMDVPANLE ARRRIAFFTN SLFMDMPRAP 1020 RVRKMLSFSV MTPYYSEETV YSKSDLEMEN EDDEWTNFME RINCKKESEV WENEENILQL 1080 RHWASLRGQT LCRTLRGMMY YRRALKLQAF LDMANESDMK LVSDFRYAEI LAGYKAITNP 1140 SEEDKKSQRS LSAQLEAVAD MKFTYVATCQ IYGNQKRKGD RHATNILNLM VKYVISGV* 1200 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
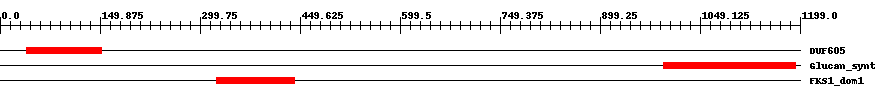
| pfam04652 | DUF605 | 2.0e-7 | 39 | 152 | 117 | + Vta1 like. Vta1 (VPS20-associated protein 1) is a positive regulator of Vps4. Vps4 is an ATPase that is required in the multivesicular body (MVB) sorting pathway to dissociate the endosomal sorting complex required for transport (ESCRT). Vta1 promotes correct assembly of Vps4 and stimulates its ATPase activity through its conserved Vta1/SBP1/LIP5 region. | ||
| pfam02364 | Glucan_synthase | 7.0e-33 | 995 | 1193 | 220 | + 1,3-beta-glucan synthase component. This family consists of various 1,3-beta-glucan synthase components including Gls1, Gls2 and Gls3 from yeast. 1,3-beta-glucan synthase EC:2.4.1.34 also known as callose synthase catalyzes the formation of a beta-1,3-glucan polymer that is a major component of the fungal cell wall. The reaction catalyzed is:- UDP-glucose + {(1,3)-beta-D-glucosyl}(N) <=> UDP + {(1,3)-beta-D-glucosyl}(N+1). | ||
| pfam14288 | FKS1_dom1 | 3.0e-52 | 325 | 441 | 118 | + 1,3-beta-glucan synthase subunit FKS1, domain-1. The FKS1_dom1 domain is likely to be the 'Class I' region just N-terminal to the first set of transmembrane helices that is involved in 1,3-beta-glucan synthesis itself. This family is found on proteins with family Glucan_synthase, pfam02364. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0000148 | 1,3-beta-D-glucan synthase complex |
| GO:0003843 | 1,3-beta-D-glucan synthase activity |
| GO:0006075 | (1->3)-beta-D-glucan biosynthetic process |
| GO:0016020 | membrane |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK49452.2 | 0 | 10 | 1191 | 26 | 1251 | AF304372_1 putative beta-1,3-glucan synthase [Nicotiana alata] |
| GenBank | ACV04900.1 | 0 | 10 | 1191 | 15 | 1238 | callose synthase 5 [Arabidopsis thaliana] |
| EMBL | CAN80181.1 | 0 | 10 | 1191 | 3 | 1253 | hypothetical protein [Vitis vinifera] |
| RefSeq | NP_849953.2 | 0 | 10 | 1191 | 15 | 1238 | CALS5 (CALLOSE SYNTHASE 5); 1,3-beta-glucan synthase [Arabidopsis thaliana] |
| RefSeq | XP_002274337.1 | 0 | 10 | 1191 | 11 | 1238 | PREDICTED: hypothetical protein [Vitis vinifera] |