

| Basic Information | |
|---|---|
| Species | Medicago truncatula |
| Cazyme ID | Medtr2g029820.1 |
| Family | AA2 |
| Protein Properties | Length: 815 Molecular Weight: 88113.6 Isoelectric Point: 8.225 |
| Chromosome | Chromosome/Scaffold: 2 Start: 10028566 End: 10035297 |
| Description | Peroxidase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| AA2 | 136 | 404 | 0 |
| VREVIRNVSKTDTRMLASLVRLHFHDCFVQGCDASVLLNNTATIVSEQDAFPNRNSLRGLDVVNQIKTAVEKACPNTVSCADILALAAELSSTLSQGPDW KVPLGRRDGLTANQSLANQNLPAPFNSLDQLKAAFASQGLSTTDLVALSGAHTFGRAHCSLFVSRLYNFSNTGSPDPTLNATYLQQLRNICPNGGPGTPL ASFDPTTPDKFDKNYYSNLQVKKGLLQSDQELFSTSGADTISIVNNFATDQKAFFESFKAAMIKMGNIG | |||
| AA2 | 504 | 772 | 0 |
| VREVIRSVSKKDPRMLGSLVRLHFHDCFVQGCDASVLLNKTDTVVSEQDAFPNRNSLRGLDVVNQIKTAVEKACPNTVSCADILALSAELSSTLADGPDW KVPLGRRDGLTANQLLANKNLPAPFNTTDQLKAAFAAQGLDTTDLVALSGAHTFGRAHCSLFVSRLYNFNGTGSPDPTLNTTYLQQLRTICPNGGPGTNL TNFDPTTPDKFDKNYYSNLQVKKGLLQSDQELFSTSGSDTISIVNKFATDQKAFFESFKAAMIKMGNIG | |||
| Full Sequence |
|---|
| Protein Sequence Length: 815 Download |
| MASISPHRKI KGVKIEKESS TQLAIGAGKA LFALMQQFQP NFNRFNEYPI VFMWLPGFGE 60 PALFSLYSAF GAMWVLGEPF MKNLKLNLTT KMNSLRAVAI ALCFIVALFG VLPFPSNAQL 120 NPSFYSKTCP NVSSIVREVI RNVSKTDTRM LASLVRLHFH DCFVQGCDAS VLLNNTATIV 180 SEQDAFPNRN SLRGLDVVNQ IKTAVEKACP NTVSCADILA LAAELSSTLS QGPDWKVPLG 240 RRDGLTANQS LANQNLPAPF NSLDQLKAAF ASQGLSTTDL VALSGAHTFG RAHCSLFVSR 300 LYNFSNTGSP DPTLNATYLQ QLRNICPNGG PGTPLASFDP TTPDKFDKNY YSNLQVKKGL 360 LQSDQELFST SGADTISIVN NFATDQKAFF ESFKAAMIKM GNIGVLTGNQ GEIRKQCNFV 420 NSKSVELGLV NVASTEEGMV MLILYTFSVS LSLSLSQQTM NSLRAVAIAL CCIVVVLGGL 480 PFSSNAQLDP SFYRNTCPNV SSIVREVIRS VSKKDPRMLG SLVRLHFHDC FVQGCDASVL 540 LNKTDTVVSE QDAFPNRNSL RGLDVVNQIK TAVEKACPNT VSCADILALS AELSSTLADG 600 PDWKVPLGRR DGLTANQLLA NKNLPAPFNT TDQLKAAFAA QGLDTTDLVA LSGAHTFGRA 660 HCSLFVSRLY NFNGTGSPDP TLNTTYLQQL RTICPNGGPG TNLTNFDPTT PDKFDKNYYS 720 NLQVKKGLLQ SDQELFSTSG SDTISIVNKF ATDQKAFFES FKAAMIKMGN IGVLTGKQGE 780 IRKQCNFVNS KSVELGLVNV ASTDSSDEGM VSSM* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
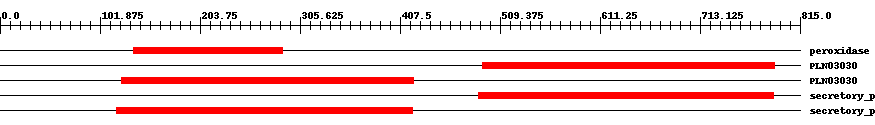 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00141 | peroxidase | 9.0e-58 | 136 | 288 | 153 | + Peroxidase. | ||
| PLN03030 | PLN03030 | 1.0e-72 | 492 | 789 | 303 | + cationic peroxidase; Provisional | ||
| PLN03030 | PLN03030 | 5.0e-79 | 124 | 421 | 303 | + cationic peroxidase; Provisional | ||
| cd00693 | secretory_peroxidase | 3.0e-163 | 487 | 788 | 302 | + Horseradish peroxidase and related secretory plant peroxidases. Secretory peroxidases belong to class III of the plant heme-dependent peroxidase superfamily. All members of the superfamily share a heme prosthetic group and catalyze a multistep oxidative reaction involving hydrogen peroxide as the electron acceptor. Class III peroxidases are found in the extracellular space or in the vacuole in plants where they have been implicated in hydrogen peroxide detoxification, auxin catabolism and lignin biosynthesis, and stress response. Class III peroxidases contain four conserved disulphide bridges and two conserved calcium binding sites. | ||
| cd00693 | secretory_peroxidase | 4.0e-170 | 119 | 420 | 302 | + Horseradish peroxidase and related secretory plant peroxidases. Secretory peroxidases belong to class III of the plant heme-dependent peroxidase superfamily. All members of the superfamily share a heme prosthetic group and catalyze a multistep oxidative reaction involving hydrogen peroxide as the electron acceptor. Class III peroxidases are found in the extracellular space or in the vacuole in plants where they have been implicated in hydrogen peroxide detoxification, auxin catabolism and lignin biosynthesis, and stress response. Class III peroxidases contain four conserved disulphide bridges and two conserved calcium binding sites. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004601 | peroxidase activity |
| GO:0006979 | response to oxidative stress |
| GO:0020037 | heme binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABO77634.1 | 0 | 92 | 440 | 1 | 353 | peroxidase [Medicago truncatula] |
| GenBank | ABO77634.1 | 0 | 460 | 814 | 1 | 356 | peroxidase [Medicago truncatula] |
| DDBJ | BAD97438.1 | 0 | 478 | 814 | 17 | 353 | peroxidase [Pisum sativum] |
| EMBL | CAA62226.1 | 0 | 92 | 440 | 1 | 352 | peroxidase1B [Medicago sativa] |
| EMBL | CAA62226.1 | 0 | 460 | 814 | 1 | 355 | peroxidase1B [Medicago sativa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1fhf_C | 0 | 487 | 789 | 1 | 303 | A Chain A, The Structure Of Soybean Peroxidase |
| PDB | 1fhf_C | 0 | 119 | 421 | 1 | 303 | A Chain A, The Structure Of Soybean Peroxidase |
| PDB | 1fhf_B | 0 | 487 | 789 | 1 | 303 | A Chain A, The Structure Of Soybean Peroxidase |
| PDB | 1fhf_B | 0 | 119 | 421 | 1 | 303 | A Chain A, The Structure Of Soybean Peroxidase |
| PDB | 1fhf_A | 0 | 487 | 789 | 1 | 303 | A Chain A, The Structure Of Soybean Peroxidase |