

| Basic Information | |
|---|---|
| Species | Cucumis sativus |
| Cazyme ID | Cucsa.333930.2 |
| Family | GH17 |
| Protein Properties | Length: 365 Molecular Weight: 39960.3 Isoelectric Point: 5.1158 |
| Chromosome | Chromosome/Scaffold: 03290 Start: 76645 End: 78677 |
| Description | Glycosyl hydrolase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 51 | 358 | 0 |
| IGVCYGTMGDNLPGAKEVVQLYKSYGIKKMRIYHPDKHILNALQGSNIQVMVGIPNSDLQSLVNNTSATNWVQTNIQAYVPNIKFQYIAVGNEVQPSDSF AAFVLPVMRNIYSAIVEANLEDQIKVSTVISANLLGSSFPPSVGSFSREANELMEPIVGFLVQNASPLLANLYPYYTYMSGTFTLDYALFNGPSVVKDGN FDYHNVFEVMVDAIYAALEKCGGTNVSIVVSESGWPSAGDGNTKVENGVAGSYYSNLIKFVQGGTQRRPGRAIETYLFAMFDENLRSPAVDKHFGLFTYD QKLKYVIN | |||
| Full Sequence |
|---|
| Protein Sequence Length: 365 Download |
| MAMAMAINNN NEHEDSMALF PFGSFNGFGS FGSGCYHPRL PSPTVAIQEV IGVCYGTMGD 60 NLPGAKEVVQ LYKSYGIKKM RIYHPDKHIL NALQGSNIQV MVGIPNSDLQ SLVNNTSATN 120 WVQTNIQAYV PNIKFQYIAV GNEVQPSDSF AAFVLPVMRN IYSAIVEANL EDQIKVSTVI 180 SANLLGSSFP PSVGSFSREA NELMEPIVGF LVQNASPLLA NLYPYYTYMS GTFTLDYALF 240 NGPSVVKDGN FDYHNVFEVM VDAIYAALEK CGGTNVSIVV SESGWPSAGD GNTKVENGVA 300 GSYYSNLIKF VQGGTQRRPG RAIETYLFAM FDENLRSPAV DKHFGLFTYD QKLKYVINLN 360 FSLY* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 2.0e-8 | 70 | 350 | 293 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 2.0e-118 | 51 | 358 | 312 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACZ74626.1 | 0 | 51 | 362 | 38 | 352 | beta-1,3-glucanase [Hevea brasiliensis] |
| RefSeq | XP_002277169.1 | 0 | 51 | 360 | 23 | 338 | PREDICTED: class I beta-1,3-glucanase [Vitis vinifera] |
| RefSeq | XP_002299791.1 | 0 | 44 | 355 | 21 | 333 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002336706.1 | 0 | 45 | 355 | 7 | 318 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002513528.1 | 0 | 49 | 357 | 31 | 340 | Lichenase precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3f55_D | 0 | 51 | 362 | 2 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| PDB | 3f55_C | 0 | 51 | 362 | 2 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| PDB | 3f55_B | 0 | 51 | 362 | 2 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| PDB | 3f55_A | 0 | 51 | 362 | 2 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| PDB | 3em5_D | 0 | 51 | 362 | 2 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO779722 | 317 | 49 | 360 | 0 |
| HS404853 | 245 | 45 | 284 | 0 |
| DY278959 | 304 | 51 | 348 | 0 |
| DY267406 | 310 | 51 | 354 | 0 |
| EL784282 | 314 | 51 | 360 | 0 |
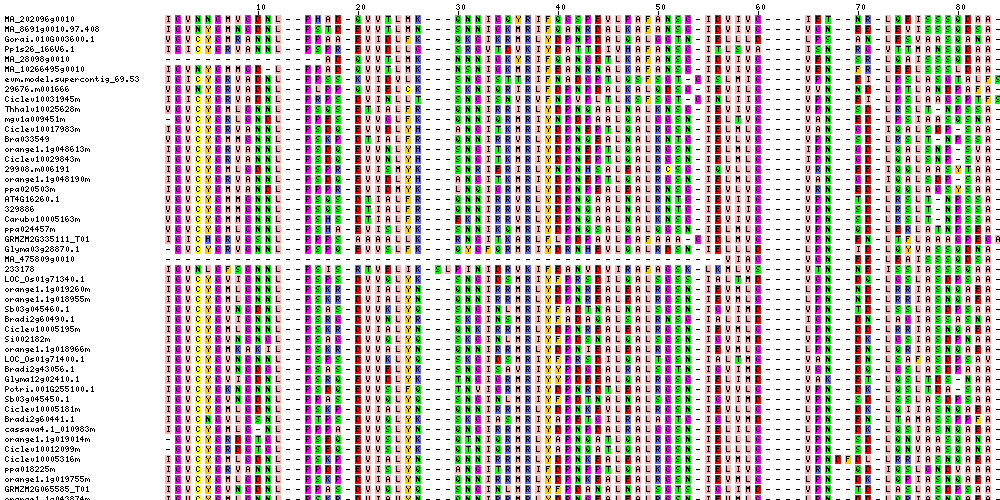
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |