

| Basic Information | |
|---|---|
| Species | Setaria italica |
| Cazyme ID | Si003802m |
| Family | GH17 |
| Protein Properties | Length: 311 Molecular Weight: 32614.2 Isoelectric Point: 5.2505 |
| Chromosome | Chromosome/Scaffold: 5 Start: 46076315 End: 46077247 |
| Description | Glycosyl hydrolase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 5 | 307 | 0 |
| GVCYGMIANNLPPPGEVVQLYKSSGIRNMRIYFPDSHVMEALSGSGIGLILGVVNQDIVGLAGCQSCAASWVQTNVRPYYPAVNILYIAVGNEVSDGAAQ SILPAMRNLQAALAAAGLAAIKVSTCVRLDVVTNTFPPSAGVFAQPYMVDIAQFLAGAGASLLANVYPYFAYRGSPGDISLNYALFLPGTTVRDGGNGLV YTNLFDAMVDAVVAALEKAGAASVRVVVSESGWPSAGGTAASVENARTYVQNLIDHAAQGTPKRPGALETFVFAMFNENQKPGELTEQNFGLFYPNKSPV YPI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 311 Download |
| VQSSGVCYGM IANNLPPPGE VVQLYKSSGI RNMRIYFPDS HVMEALSGSG IGLILGVVNQ 60 DIVGLAGCQS CAASWVQTNV RPYYPAVNIL YIAVGNEVSD GAAQSILPAM RNLQAALAAA 120 GLAAIKVSTC VRLDVVTNTF PPSAGVFAQP YMVDIAQFLA GAGASLLANV YPYFAYRGSP 180 GDISLNYALF LPGTTVRDGG NGLVYTNLFD AMVDAVVAAL EKAGAASVRV VVSESGWPSA 240 GGTAASVENA RTYVQNLIDH AAQGTPKRPG ALETFVFAMF NENQKPGELT EQNFGLFYPN 300 KSPVYPIIFR * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 0.0003 | 230 | 301 | 78 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 6.0e-136 | 5 | 309 | 311 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAB63854.1 | 0 | 5 | 310 | 32 | 337 | putative beta 1,3-glucanase [Oryza sativa Japonica Group] |
| RefSeq | NP_001045372.1 | 0 | 1 | 309 | 27 | 334 | Os01g0944700 [Oryza sativa (japonica cultivar-group)] |
| RefSeq | NP_001045373.1 | 0 | 1 | 310 | 6 | 315 | Os01g0944800 [Oryza sativa (japonica cultivar-group)] |
| RefSeq | XP_002459077.1 | 0 | 1 | 309 | 27 | 336 | hypothetical protein SORBIDRAFT_03g045490 [Sorghum bicolor] |
| RefSeq | XP_002459079.1 | 0 | 1 | 310 | 28 | 340 | hypothetical protein SORBIDRAFT_03g045510 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1ghs_B | 0 | 5 | 309 | 2 | 306 | A Chain A, The Three-Dimensional Structures Of Two Plant Beta-Glucan Endohydrolases With Distinct Substrate Specificities |
| PDB | 1ghs_A | 0 | 5 | 309 | 2 | 306 | A Chain A, The Three-Dimensional Structures Of Two Plant Beta-Glucan Endohydrolases With Distinct Substrate Specificities |
| PDB | 2cyg_A | 0 | 5 | 309 | 2 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_D | 0 | 5 | 309 | 3 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_C | 0 | 5 | 309 | 3 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DR739897 | 304 | 5 | 307 | 0 |
| DR739798 | 309 | 5 | 309 | 0 |
| HX143290 | 240 | 72 | 311 | 0 |
| BU103698 | 312 | 1 | 311 | 0 |
| GO874911 | 302 | 1 | 300 | 0 |
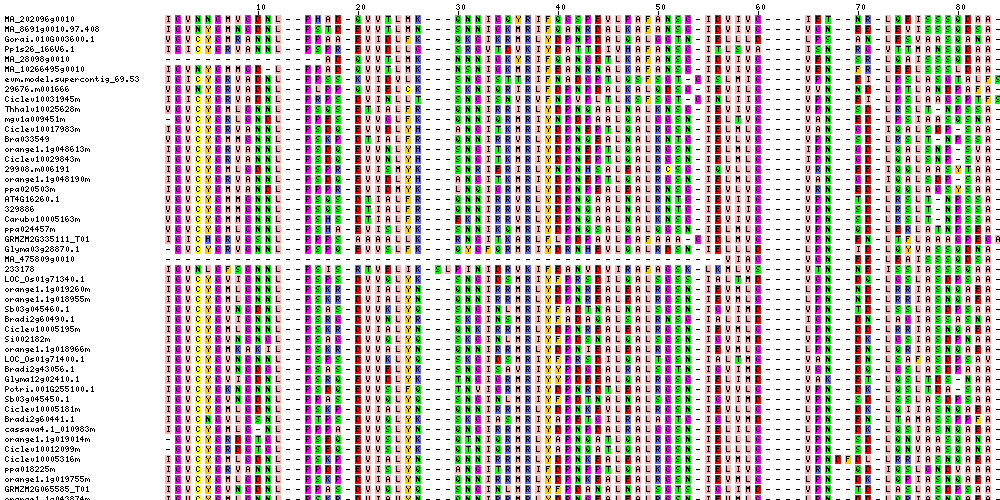
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |