

| Basic Information | |
|---|---|
| Species | Linum usitatissimum |
| Cazyme ID | Lus10031033 |
| Family | GH17 |
| Protein Properties | Length: 197 Molecular Weight: 22032.1 Isoelectric Point: 5.8561 |
| Chromosome | Chromosome/Scaffold: 261 Start: 715006 End: 716181 |
| Description | beta-1,3-glucanase 1 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 2 | 195 | 0 |
| QRSITAIGLNYRKVSTTIRYSLKKSFPPSKGAFTAEVDVLMRPLVKVLKRNGAPLMVNSHPYYGTPREYVFFGEGYEGVKDGEYRYTNMFDAQVDAMYAA LERYEGGEAVKVVVAETGWPTAGDLDGMASVENAKEFNGNLAKHVKLGTPRRPGEEIETYVFELFNENMKTPVGTENCWGVFYADMTPVYPMTF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 197 Download |
| MQRSITAIGL NYRKVSTTIR YSLKKSFPPS KGAFTAEVDV LMRPLVKVLK RNGAPLMVNS 60 HPYYGTPREY VFFGEGYEGV KDGEYRYTNM FDAQVDAMYA ALERYEGGEA VKVVVAETGW 120 PTAGDLDGMA SVENAKEFNG NLAKHVKLGT PRRPGEEIET YVFELFNENM KTPVGTENCW 180 GVFYADMTPV YPMTFQ* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 5.0e-7 | 118 | 186 | 73 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 7.0e-57 | 2 | 195 | 203 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABD85024.1 | 0 | 1 | 195 | 138 | 336 | beta-1,3-glucanase [Lilium hybrid division VII] |
| GenBank | ACF06549.1 | 0 | 4 | 195 | 142 | 338 | beta-1,3-glucanase [Elaeis guineensis] |
| GenBank | ACF06550.1 | 0 | 4 | 195 | 142 | 338 | beta-1,3-glucanase [Elaeis guineensis] |
| RefSeq | XP_002275039.1 | 0 | 1 | 191 | 145 | 343 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002533898.1 | 0 | 2 | 196 | 59 | 257 | Glucan endo-1,3-beta-glucosidase, basic isoform precursor, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 4 | 195 | 115 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1ghr_A | 1.4013e-45 | 1 | 195 | 109 | 306 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1aq0_B | 1.4013e-45 | 1 | 195 | 109 | 306 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| PDB | 1aq0_A | 1.4013e-45 | 1 | 195 | 109 | 306 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| PDB | 3f55_D | 1e-39 | 1 | 196 | 116 | 316 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EY400440 | 202 | 3 | 195 | 0 |
| EY409234 | 190 | 14 | 195 | 0 |
| EY399346 | 202 | 3 | 195 | 0 |
| EY400962 | 180 | 23 | 195 | 1.4013e-45 |
| EY404571 | 203 | 3 | 195 | 4.06377e-44 |
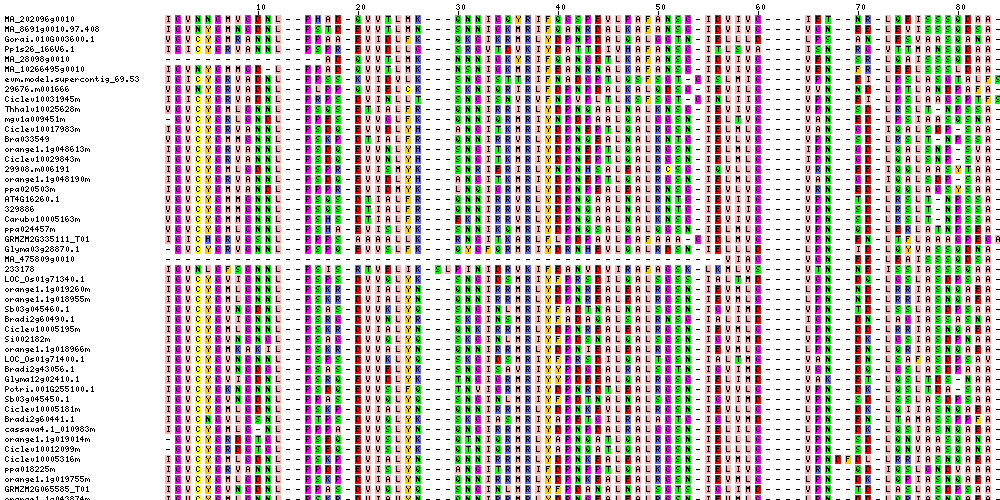
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |