

| Basic Information | |
|---|---|
| Species | Linum usitatissimum |
| Cazyme ID | Lus10016883 |
| Family | GH17 |
| Protein Properties | Length: 331 Molecular Weight: 37081.9 Isoelectric Point: 4.9684 |
| Chromosome | Chromosome/Scaffold: 153 Start: 429213 End: 431102 |
| Description | Glycosyl hydrolase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 1 | 306 | 0 |
| MGNNLPPPSEVVSLCQQNNIWRMRLYDPNRDALWALRDSGIEVTIGVPNSDLKHLNNWDDAYWWVQEYVRNNWPNVKVKYIAVGNEVSPMYNADLASAVL PAMRNIYNALVQMGLHEQVKVSTAIDMTLLANSYPPSAGAFRDDIRWFLDPIIGFLGSVKAPLLANIYTYFSYRDNPSTISLPYALLSSQSVSVWDNGLG YTNLFDAMLDSLYSAVERLGGWSVEVVVSESGWPSAGAGAATTMENARVFYTNLVQQVKRGSPKRPNKAIETYLFAMFDENQKDPELERNFGVFYPYKQS KYNLNF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 331 Download |
| MGNNLPPPSE VVSLCQQNNI WRMRLYDPNR DALWALRDSG IEVTIGVPNS DLKHLNNWDD 60 AYWWVQEYVR NNWPNVKVKY IAVGNEVSPM YNADLASAVL PAMRNIYNAL VQMGLHEQVK 120 VSTAIDMTLL ANSYPPSAGA FRDDIRWFLD PIIGFLGSVK APLLANIYTY FSYRDNPSTI 180 SLPYALLSSQ SVSVWDNGLG YTNLFDAMLD SLYSAVERLG GWSVEVVVSE SGWPSAGAGA 240 ATTMENARVF YTNLVQQVKR GSPKRPNKAI ETYLFAMFDE NQKDPELERN FGVFYPYKQS 300 KYNLNFGAAR DTWDVSSQLL HNATTTVSAI * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00332 | Glyco_hydro_17 | 2.0e-130 | 1 | 306 | 308 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAA87456.1 | 0 | 2 | 324 | 46 | 366 | beta-1,3-glucanase [Hevea brasiliensis] |
| GenBank | AAP87281.1 | 0 | 2 | 324 | 46 | 366 | beta-1,3-glucanase [Hevea brasiliensis] |
| GenBank | ABG49448.1 | 0 | 2 | 324 | 46 | 366 | beta-1,3-glucanase [Hevea brasiliensis] |
| GenBank | ACZ74626.1 | 0 | 1 | 324 | 45 | 366 | beta-1,3-glucanase [Hevea brasiliensis] |
| Swiss-Prot | P52407 | 0 | 2 | 324 | 46 | 366 | E13B_HEVBR RecName: Full=Glucan endo-1,3-beta-glucosidase, basic vacuolar isoform; AltName: Full=(1- |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3f55_D | 0 | 2 | 306 | 10 | 315 | A Chain A, Crystal Structure Of Class I Chitinase From Oryza Sativa L. Japonica |
| PDB | 3f55_C | 0 | 2 | 306 | 10 | 315 | A Chain A, Crystal Structure Of Class I Chitinase From Oryza Sativa L. Japonica |
| PDB | 3f55_B | 0 | 2 | 306 | 10 | 315 | A Chain A, Crystal Structure Of Class I Chitinase From Oryza Sativa L. Japonica |
| PDB | 3f55_A | 0 | 2 | 306 | 10 | 315 | A Chain A, Crystal Structure Of Class I Chitinase From Oryza Sativa L. Japonica |
| PDB | 3em5_D | 0 | 2 | 306 | 10 | 315 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
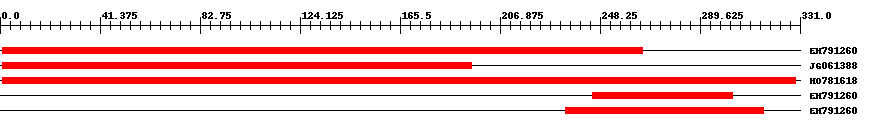 | ||||
| Hit | Length | Start | End | EValue |
| EH791260 | 266 | 1 | 266 | 0 |
| JG061388 | 195 | 1 | 195 | 0 |
| HO781618 | 331 | 1 | 329 | 0 |
| EH791260 | 59 | 245 | 303 | 0.0000003 |
| EH791260 | 84 | 234 | 316 | 0.000009 |
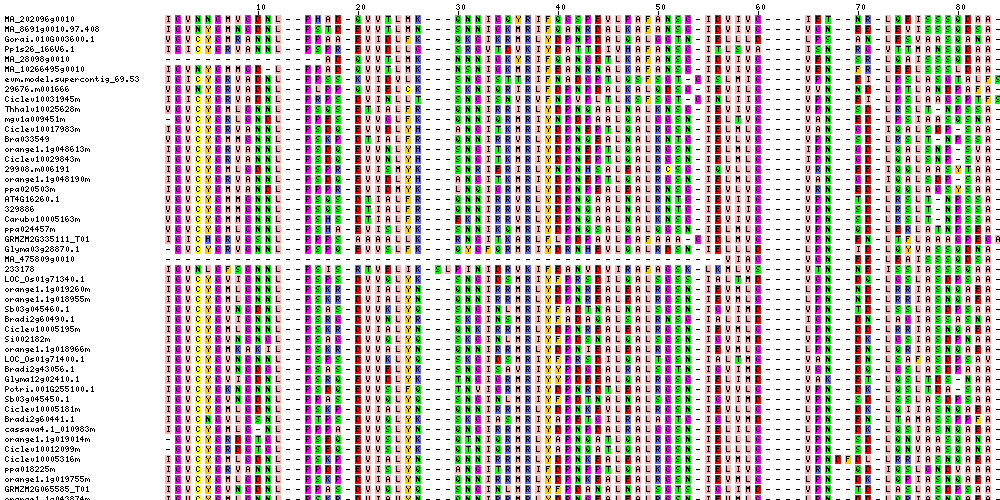
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |