

| Basic Information | |
|---|---|
| Species | Linum usitatissimum |
| Cazyme ID | Lus10027479 |
| Family | GH17 |
| Protein Properties | Length: 271 Molecular Weight: 29611.3 Isoelectric Point: 4.5147 |
| Chromosome | Chromosome/Scaffold: 96 Start: 564055 End: 564867 |
| Description | Glycosyl hydrolase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 1 | 265 | 0 |
| MEVVLGTVNSNLEQLATNPSFAAQWVSANVVPYHPALKLRYVSAGNEVIPGPLAQHVLGATENLDNALKSSGIDNVCVSTATSFVSIGESFPPSAGEFAD ESKPFMAPIIEFLAENDYPLLVNVYPYFAYKSNPDRIDLDYALGNPGRIFVADGRLRYRSLFDAMVDAMYAAAEKVHVGKVRMVVTETGWPSAGDRVATM KNAKAYVNNAAARAGGSDAGTPRRPGIETETYLFALFNEDLKPEGEERNFGMFWPDFTEVYHVDF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 271 Download |
| MEVVLGTVNS NLEQLATNPS FAAQWVSANV VPYHPALKLR YVSAGNEVIP GPLAQHVLGA 60 TENLDNALKS SGIDNVCVST ATSFVSIGES FPPSAGEFAD ESKPFMAPII EFLAENDYPL 120 LVNVYPYFAY KSNPDRIDLD YALGNPGRIF VADGRLRYRS LFDAMVDAMY AAAEKVHVGK 180 VRMVVTETGW PSAGDRVATM KNAKAYVNNA AARAGGSDAG TPRRPGIETE TYLFALFNED 240 LKPEGEERNF GMFWPDFTEV YHVDFPQYFN * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00332 | Glyco_hydro_17 | 1.0e-91 | 2 | 265 | 266 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| DDBJ | BAA77786.1 | 0 | 3 | 265 | 48 | 312 | beta-1,3-glucanase [Oryza sativa] |
| EMBL | CBI33865.1 | 0 | 1 | 265 | 75 | 340 | unnamed protein product [Vitis vinifera] |
| GenBank | EAY98549.1 | 0 | 1 | 265 | 86 | 352 | hypothetical protein OsI_20461 [Oryza sativa Indica Group] |
| RefSeq | NP_001055934.1 | 0 | 1 | 265 | 80 | 346 | Os05g0495900 [Oryza sativa (japonica cultivar-group)] |
| RefSeq | XP_002277609.1 | 0 | 1 | 265 | 63 | 328 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 1 | 265 | 48 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_D | 0 | 1 | 265 | 49 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_C | 0 | 1 | 265 | 49 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_B | 0 | 1 | 265 | 49 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_A | 0 | 1 | 265 | 49 | 315 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| EY400440 | 227 | 41 | 265 | 0 |
| EY399346 | 228 | 40 | 265 | 0 |
| FE656899 | 231 | 8 | 235 | 0 |
| GR143023 | 221 | 46 | 265 | 0 |
| DR097437 | 270 | 1 | 265 | 0 |
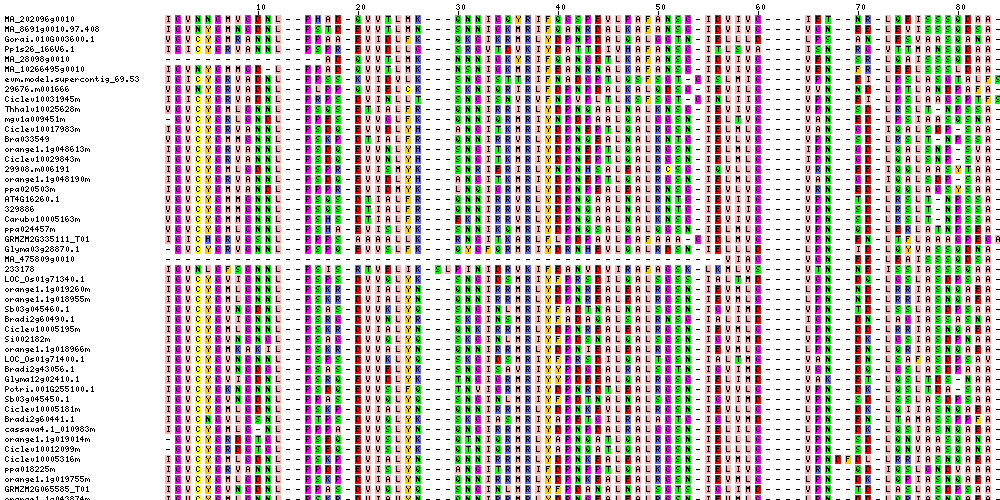
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |