

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_10432716g0030 |
| Family | GH17 |
| Protein Properties | Length: 263 Molecular Weight: 28878.6 Isoelectric Point: 7.6551 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 57 | 261 | 0 |
| RRNIQTAIQNANLQNNIKVFTTHASDVSNGFPPSRGVFKDEVKDTMRSLLQFLSDHGSPFPANIYPYFSYTGNMGSISLDYALFKSTSTVVQDGGRSYNN LFDALVDTLLSAMEALGYPNIPLIITESGWPSGGADVATVDNARAYNNNLIRHVLSNAGTPKRPGTSIETYIFALFNENKKTGPETERNFGLFNPNQQSV YSVKF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 263 Download |
| MQPVACGTAS NLGKLALPNI FFPLMDNTVG LTITFPHSIP LQISNILQWA PRFGQERRNI 60 QTAIQNANLQ NNIKVFTTHA SDVSNGFPPS RGVFKDEVKD TMRSLLQFLS DHGSPFPANI 120 YPYFSYTGNM GSISLDYALF KSTSTVVQDG GRSYNNLFDA LVDTLLSAME ALGYPNIPLI 180 ITESGWPSGG ADVATVDNAR AYNNNLIRHV LSNAGTPKRP GTSIETYIFA LFNENKKTGP 240 ETERNFGLFN PNQQSVYSVK FSP |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 0.005 | 176 | 252 | 83 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 4.0e-83 | 58 | 261 | 205 | + Glycosyl hydrolases family 17. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAV66572.1 | 0 | 59 | 263 | 141 | 343 | glucanase-like protein [Thuja occidentalis] |
| GenBank | ABB04448.1 | 0 | 58 | 262 | 136 | 334 | pathogenesis-related protein 6 [Zea mays subsp. parviglumis] |
| GenBank | ABK23947.1 | 0 | 59 | 263 | 140 | 344 | unknown [Picea sitchensis] |
| GenBank | ABK25991.1 | 0 | 51 | 263 | 128 | 342 | unknown [Picea sitchensis] |
| DDBJ | BAD93486.1 | 0 | 58 | 262 | 143 | 346 | pollen allergen CJP38 [Cryptomeria japonica] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3f55_D | 0 | 58 | 262 | 114 | 316 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_C | 0 | 58 | 262 | 114 | 316 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_B | 0 | 58 | 262 | 114 | 316 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3f55_A | 0 | 58 | 262 | 114 | 316 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 3em5_D | 0 | 58 | 262 | 114 | 316 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| ES255879 | 206 | 58 | 263 | 0 |
| EX430725 | 214 | 48 | 261 | 0 |
| CO483293 | 214 | 48 | 261 | 0 |
| CO486397 | 203 | 59 | 261 | 0 |
| CO483293 | 32 | 26 | 57 | 0.002 |
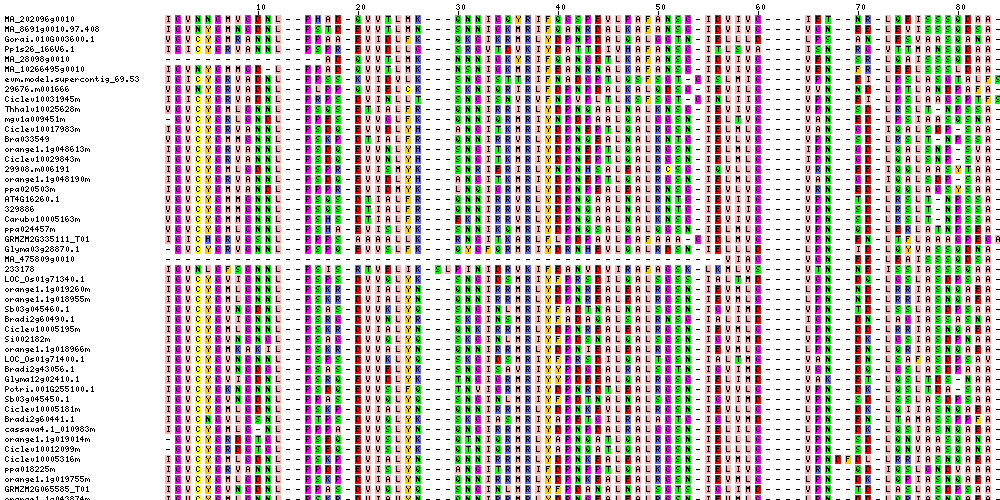
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |