

| Basic Information | |
|---|---|
| Species | Linum usitatissimum |
| Cazyme ID | Lus10031035 |
| Family | GH17 |
| Protein Properties | Length: 326 Molecular Weight: 36321.2 Isoelectric Point: 6.626 |
| Chromosome | Chromosome/Scaffold: 261 Start: 719430 End: 720547 |
| Description | beta-1,3-glucanase 2 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 44 | 324 | 0 |
| RSEPEVLRALQGTNIQLSLDITADNIPNLASSQAAADSYICDHIQPYIGGVKLRYIAIGNEVEPGQPSFQYIVLAIENIQRSISAIGFNDTKVTTCFKYN LNKTIPPSAGSFSLESESVLRPLVNVLKTNGAQYMVNVYPFFAYHKDPQKVGIREFALFMSKSPVMRDGEYEYSNLFDATVDAVYSALEKYGGASVKVVV PETGWATVGELAEVTSVENARLYNSNLVKHVYGGKGTPKRAGEEIEAYIFELFNENQKSVGIESNFGLFYPDQRPVYPMSF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 326 Download |
| MFNVDACRCG HRCVLWKQRR QPPISTRSNI TLPKIQHHHD TDIRSEPEVL RALQGTNIQL 60 SLDITADNIP NLASSQAAAD SYICDHIQPY IGGVKLRYIA IGNEVEPGQP SFQYIVLAIE 120 NIQRSISAIG FNDTKVTTCF KYNLNKTIPP SAGSFSLESE SVLRPLVNVL KTNGAQYMVN 180 VYPFFAYHKD PQKVGIREFA LFMSKSPVMR DGEYEYSNLF DATVDAVYSA LEKYGGASVK 240 VVVPETGWAT VGELAEVTSV ENARLYNSNL VKHVYGGKGT PKRAGEEIEA YIFELFNENQ 300 KSVGIESNFG LFYPDQRPVY PMSFL* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 0.001 | 236 | 315 | 85 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 1.0e-86 | 45 | 324 | 283 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABD85024.1 | 0 | 44 | 324 | 62 | 336 | beta-1,3-glucanase [Lilium hybrid division VII] |
| GenBank | ABJ98942.1 | 0 | 50 | 324 | 68 | 340 | beta-1,3-glucanase [Musa x paradisiaca] |
| GenBank | ACF06549.1 | 0 | 50 | 324 | 68 | 338 | beta-1,3-glucanase [Elaeis guineensis] |
| GenBank | ACF06550.1 | 0 | 50 | 324 | 68 | 338 | beta-1,3-glucanase [Elaeis guineensis] |
| GenBank | ACP43630.2 | 0 | 24 | 324 | 7 | 304 | beta-1,3-glucanase [Musa AB Group] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 50 | 324 | 40 | 312 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1ghr_A | 0 | 50 | 324 | 40 | 306 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1aq0_B | 0 | 50 | 324 | 40 | 306 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| PDB | 1aq0_A | 0 | 50 | 324 | 40 | 306 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| PDB | 3f55_D | 0 | 22 | 324 | 14 | 315 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DV702548 | 281 | 49 | 324 | 0 |
| EY400440 | 229 | 98 | 324 | 0 |
| EY399346 | 230 | 97 | 324 | 0 |
| DV703316 | 272 | 49 | 315 | 0 |
| DV713096 | 282 | 49 | 325 | 0 |
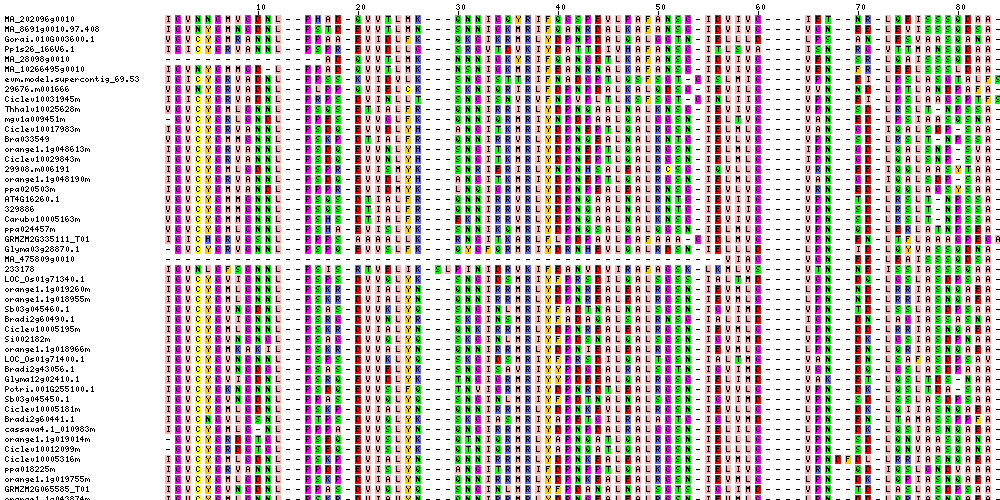
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |