

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000272624 |
| Family | GT14 |
| Protein Properties | Length: 1123 Molecular Weight: 125607 Isoelectric Point: 6.2275 |
| Chromosome | Chromosome/Scaffold: 003068193 Start: 4797 End: 16430 |
| Description | AAA-ATPase 1 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT14 | 914 | 1032 | 1.79366e-43 |
| DQRAKPVIIDPGLYSRQKSDVFWISQKRSVPTAYKLFTGSAWMMLSRPFIEYILWGWDNLPRTVLMYYANFLSSPEGYFHTVICNAEEFQNTTVNHDLHF ISWDNPPKQHPHFLVMDDF | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1123 Download |
| MSASAWAILR ECFPESVLRH INWIYPVVAS SIYNPYIQIK FSERAGEHSL MHSEAFSAID 60 NYLRSKVSKE ARRMCGYVKK NQSLVVSMDH NEEVADEFQG AKVWWALEKT IDMQRSAHRE 120 RWYYKLTFHR THRALIVGPY IKHVMKEGNA TKMGNRERKL YTNNGARWSH VVFEHPATFK 180 TLAMDPVKKQ EIIDDLMAFS KAEEFYKSTG RAWKRGYLLY GPPGTGKSTM IAAMANLLGY 240 DLYDLELTSV KNNTELRKLL LETSNKSVIV IEDIDCSLDL TGQRSKKKYK VIEDEDTGDG 300 KEKEGEEKEA NIPSQVTLSG LLNFIDGLWS ACKGERLIVF TTNYVEKLDA ALTRKGRLDK 360 HIELSYCTFE AFKVLARNYL KLESHHLFPA ICELLGETNV VSADVAEHLM RKTLSRNVDI 420 CLQNLIQALE KEKENGKLIA EEEGKDGNNQ LKRNSCWWVL VTLSCPPGQL DFFITAPTGV 480 KTTSTGLFLS ALSFGFFFSS LXVWXVNKVT GGKNGGWLAK TINQGRLDCF YGLLTILVVG 540 EFGILGFYTH FSSCILPGIE VQGNLDQNQP CGLERHGQQL QCDSPRVEFG LPVRRDGDAD 600 GDGDHVEHGV ALESVLLEGD ADGVDGDGHE GLEHLDEGDG EEDVGXVGEP EGEGVEGADG 660 DDRLEVCXLR RLRSWAKKKT ELGRSQSRAK IQISCQPVNE TDHYWVAVKD TLTNRIVISL 720 VAGDNGYQXA TEEEEMVFTL GGFTSSLHPA NLHYCFYLFQ LVFRNILPTP CEKTPPTFRG 780 IEAKDFGNFX KPSSQIGLLD LRLRRRRRQL EADAESAVPS EESVRSAFGP GGVGGGAAGL 840 GEICEGGATV CQAGECQGXG EGQFGDVQRA DDGHQYAARG GDFVEGGRRV GLKWVRSVXE 900 NNVYALSLPR KLGDQRAKPV IIDPGLYSRQ KSDVFWISQK RSVPTAYKLF TGSAWMMLSR 960 PFIEYILWGW DNLPRTVLMY YANFLSSPEG YFHTVICNAE EFQNTTVNHD LHFISWDNPP 1020 KQHPHFLVMD DFQRMVDSNA PFARKFGRNE SVLDKIDSEL LGCNADGYVP GGWFNTQANG 1080 SATVPDYSLK NITTLRPGVG AQRLKHLVTS LISAEDFHAK QCT |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
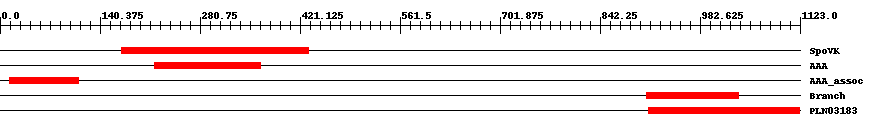 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0464 | SpoVK | 7.0e-11 | 171 | 433 | 278 | + ATPases of the AAA+ class [Posttranslational modification, protein turnover, chaperones] | ||
| pfam00004 | AAA | 8.0e-18 | 217 | 366 | 159 | + ATPase family associated with various cellular activities (AAA). AAA family proteins often perform chaperone-like functions that assist in the assembly, operation, or disassembly of protein complexes. | ||
| pfam14363 | AAA_assoc | 4.0e-20 | 13 | 110 | 102 | + Domain associated at C-terminal with AAA. This domain is found in association with the AAA family, pfam00004. | ||
| pfam02485 | Branch | 1.0e-26 | 907 | 1036 | 134 | + Core-2/I-Branching enzyme. This is a family of two different beta-1,6-N-acetylglucosaminyltransferase enzymes, I-branching enzyme and core-2 branching enzyme . I-branching enzyme is responsible for the production of the blood group I-antigen during embryonic development. Core-2 branching enzyme forms crucial side-chain branches in O-glycans. | ||
| PLN03183 | PLN03183 | 4.0e-97 | 911 | 1122 | 215 | + acetylglucosaminyltransferase family protein; Provisional | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0005524 | ATP binding |
| GO:0008375 | acetylglucosaminyltransferase activity |
| GO:0016020 | membrane |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACU20008.1 | 0 | 51 | 418 | 1 | 373 | unknown [Glycine max] |
| EMBL | CAN77016.1 | 0 | 6 | 438 | 19 | 457 | hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002280981.1 | 0 | 5 | 444 | 69 | 509 | PREDICTED: hypothetical protein isoform 1 [Vitis vinifera] |
| RefSeq | XP_002319660.1 | 0 | 6 | 425 | 16 | 439 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002325775.1 | 0 | 5 | 429 | 15 | 441 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2r65_E | 0.00000004 | 179 | 362 | 10 | 178 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2r65_D | 0.00000004 | 179 | 362 | 10 | 178 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2r65_C | 0.00000004 | 179 | 362 | 10 | 178 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2r65_B | 0.00000004 | 179 | 362 | 10 | 178 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |
| PDB | 2r65_A | 0.00000004 | 179 | 362 | 10 | 178 | A Chain A, Crystal Structure Of Medicago Truncatula Ugt85h2- Insights Into The Structural Basis Of A Multifunctional (Iso) Flavonoid Glycosyltransferase |