

| Basic Information | |
|---|---|
| Species | Populus trichocarpa |
| Cazyme ID | Potri.007G018600.3 |
| Family | AA4 |
| Protein Properties | Length: 436 Molecular Weight: 47512.6 Isoelectric Point: 6.1917 |
| Chromosome | Chromosome/Scaffold: 07 Start: 1430882 End: 1437643 |
| Description | FAD-linked oxidases family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
| Family | Start | End | Evalue |
| AA4 | 10 | 179 | 1.4e-21 |
| LLLLPRNTEEVSKILEYCNSRRLAVVPQGGNTGLVGGSVPVFDEVIINAGSMNKIIAFDKVSGILVCEAGCILENLISYLDNQGFIMPLDLGAKGSCQIG GNVSTNAGGLRFVRYGSLHGNVLGLEAVLANGDVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIVTKVSIL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 436 Download |
| MHKYKGSSKL LLLPRNTEEV SKILEYCNSR RLAVVPQGGN TGLVGGSVPV FDEVIINAGS 60 MNKIIAFDKV SGILVCEAGC ILENLISYLD NQGFIMPLDL GAKGSCQIGG NVSTNAGGLR 120 FVRYGSLHGN VLGLEAVLAN GDVLDMLGTL RKDNTGYDLK HLFIGSEGSL GIVTKVSILT 180 PPKLSSVNIA FLACEDYLSC QKLLSEAKRK LGEILSAFEF LDSHAMDLVL NHLEGVRNPL 240 PSAVHNFYVL IETTGSDESY DKEKLEAFLL HSMESGLISD GVLAQDINQA SSFWRIREGV 300 PEALMRAGPV YKYDLSIPVE KMYSLVEDMR LRLGQSANVV GYGHLGDGNL HLNISAPRYD 360 DTILAQIEPY VYEWTSKHRG SISAEHGLGL MKANEIFYSK SHETVQLMAS IKKLLDPNGI 420 LNPYKVLPHS LCSYS* |
| Functional Domains Download unfiltered results here | ||||||
|---|---|---|---|---|---|---|
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description |
| PRK11230 | PRK11230 | 4.0e-39 | 4 | 428 | 451 | + glycolate oxidase subunit GlcD; Provisional |
| PLN02805 | PLN02805 | 1.0e-43 | 2 | 432 | 444 | + D-lactate dehydrogenase [cytochrome] |
| pfam02913 | FAD-oxidase_C | 9.0e-61 | 183 | 426 | 252 | + FAD linked oxidases, C-terminal domain. This domain has a ferredoxin-like fold. |
| TIGR00387 | glcD | 1.0e-78 | 12 | 425 | 429 | + glycolate oxidase, subunit GlcD. This protein, the glycolate oxidase GlcD subunit, is similar in sequence to that of several D-lactate dehydrogenases, including that of E. coli. The glycolate oxidase has been found to have some D-lactate dehydrogenase activity [Energy metabolism, Other]. |
| COG0277 | GlcD | 3.0e-105 | 2 | 429 | 439 | + FAD/FMN-containing dehydrogenases [Energy production and conversion] |
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0008762 | UDP-N-acetylmuramate dehydrogenase activity |
| GO:0016491 | oxidoreductase activity |
| GO:0050660 | flavin adenine dinucleotide binding |
| GO:0055114 | oxidation-reduction process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI17437.1 | 0 | 1 | 435 | 1 | 435 | unnamed protein product [Vitis vinifera] |
| RefSeq | NP_568003.2 | 0 | 1 | 433 | 127 | 558 | FAD linked oxidase family protein [Arabidopsis thaliana] |
| RefSeq | XP_002268002.1 | 0 | 1 | 435 | 118 | 552 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310828.1 | 0 | 1 | 435 | 99 | 530 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002521506.1 | 0 | 1 | 434 | 133 | 565 | d-lactate dehydrognease 2, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3pm9_F | 0 | 4 | 427 | 48 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_E | 0 | 4 | 427 | 48 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_D | 0 | 4 | 427 | 48 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_C | 0 | 4 | 427 | 48 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| PDB | 3pm9_B | 0 | 4 | 427 | 48 | 476 | A Chain A, Crystal Structure Of A Putative Dehydrogenase (Rpa1076) From Rhodopseudomonas Palustris Cga009 At 2.57 A Resolution |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
| Hit | Length | Start | End | EValue |
| GO838775 | 330 | 14 | 343 | 0 |
| GO839525 | 328 | 3 | 330 | 0 |
| CO071824 | 280 | 1 | 280 | 0 |
| CO071363 | 279 | 1 | 279 | 0 |
| HO419275 | 401 | 31 | 430 | 0 |
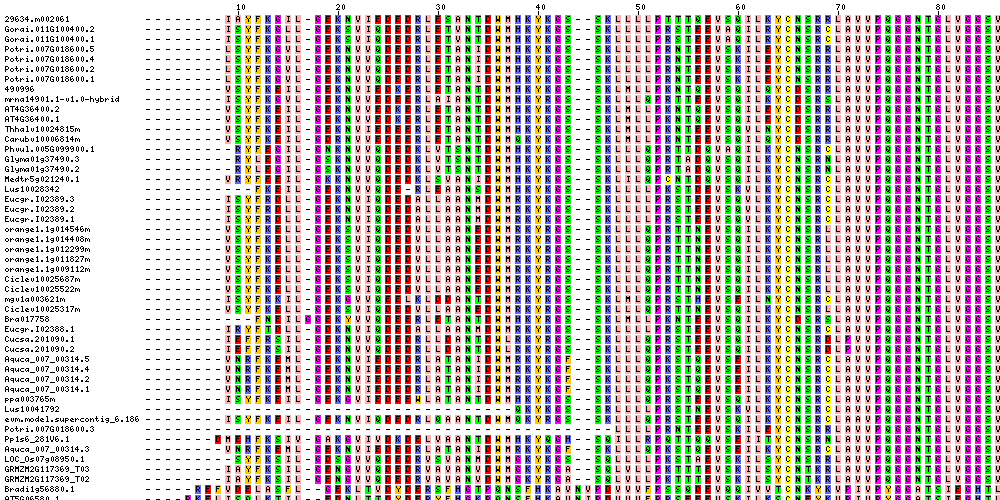
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |