

| Basic Information | |
|---|---|
| Species | Glycine max |
| Cazyme ID | Glyma18g01570.2 |
| Family | PL1 |
| Protein Properties | Length: 393 Molecular Weight: 43268.3 Isoelectric Point: 9.2666 |
| Chromosome | Chromosome/Scaffold: 18 Start: 867982 End: 870277 |
| Description | Pectate lyase family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| PL1 | 120 | 311 | 0 |
| LWIIFKHDMVIKLHKDLLVNSYKTIDGRGATIHIAGGGPCIRVQKKTNIIIHGIHIHDCKRGGVDIARSDGDGITIFGGSHVWVDHCSLSNCFDGLIDVV HGSTAITISNNNMTHHNKVMLLGHSDSYKADKNMQVTIAFNHFGVGLGGRMPRCRFGYFHVVNNDYTNWQHYAIGGSSSPTIFSQGNRFRAP | |||
| Full Sequence |
|---|
| Protein Sequence Length: 393 Download |
| MAFSFTFMFQ FLLLAPSVIY ASPVQDPELV IQEVQKSING SRRNLGYLSC GTGNPIDDCW 60 RCDPNWERNR KRLASCAIGF GKHAIGGKDG KIYVVTDPSD NPVNPKPGTL RHGVIQQEPL 120 WIIFKHDMVI KLHKDLLVNS YKTIDGRGAT IHIAGGGPCI RVQKKTNIII HGIHIHDCKR 180 GGVDIARSDG DGITIFGGSH VWVDHCSLSN CFDGLIDVVH GSTAITISNN NMTHHNKVML 240 LGHSDSYKAD KNMQVTIAFN HFGVGLGGRM PRCRFGYFHV VNNDYTNWQH YAIGGSSSPT 300 IFSQGNRFRA PNDEDHKEVT KHFKSSKSEW RKWNWRSEGD LMLNGAFFTA SGAGATARYD 360 KASSMAARPP MLVVSMTAGA GALRCNKGNL CH* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
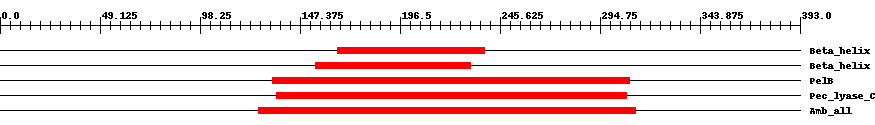 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam13229 | Beta_helix | 0.002 | 166 | 238 | 86 | + Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. | ||
| pfam13229 | Beta_helix | 0.002 | 155 | 231 | 91 | + Right handed beta helix region. This region contains a parallel beta helix region that shares some similarity with Pectate lyases. | ||
| COG3866 | PelB | 3.0e-28 | 134 | 309 | 188 | + Pectate lyase [Carbohydrate transport and metabolism] | ||
| pfam00544 | Pec_lyase_C | 5.0e-67 | 136 | 308 | 199 | + Pectate lyase. This enzyme forms a right handed beta helix structure. Pectate lyase is an enzyme involved in the maceration and soft rotting of plant tissue. | ||
| smart00656 | Amb_all | 2.0e-78 | 127 | 312 | 197 | + Amb_all domain. | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABV32549.1 | 0 | 15 | 391 | 23 | 413 | pectase lyase [Prunus persica] |
| EMBL | CBI14899.1 | 0 | 16 | 391 | 17 | 403 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002275781.1 | 0 | 16 | 391 | 72 | 458 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002285340.1 | 0 | 21 | 391 | 22 | 403 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002322215.1 | 0 | 15 | 391 | 16 | 403 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1pxz_B | 0 | 54 | 352 | 2 | 313 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 1pxz_A | 0 | 54 | 352 | 2 | 313 | A Chain A, 1.7 Angstrom Crystal Structure Of Jun A 1, The Major Allergen From Cedar Pollen |
| PDB | 3zsc_A | 7e-17 | 122 | 299 | 50 | 227 | A Chain A, Catalytic Function And Substrate Recognition Of The Pectate Lyase From Thermotoga Maritima |
| PDB | 1vbl_A | 2e-16 | 136 | 321 | 128 | 343 | A Chain A, Structure Of The Thermostable Pectate Lyase Pl 47 |
| PDB | 1pe9_B | 0.000000000000007 | 167 | 382 | 115 | 358 | A Chain A, Structure Of The Thermostable Pectate Lyase Pl 47 |