

| Basic Information | |
|---|---|
| Species | Arabidopsis lyrata |
| Cazyme ID | 489289 |
| Family | GT5 |
| Protein Properties | Length: 656 Molecular Weight: 72227.7 Isoelectric Point: 5.9188 |
| Chromosome | Chromosome/Scaffold: 6 Start: 10541641 End: 10545886 |
| Description | Glycogen/starch synthases, ADP-glucose type |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 147 | 645 | 0 |
| LVFVTSEAAPYSKTGGLGDVCGSLPIALAGRGHRVMVVSPRYLNGTAADNNYARAKDLGIRITVNCFGGSQEVSFYHEYRDGVDWVFVDHKSYHRPGNPY GDSKGAFGDNQFRFTLLCHAACEAPLVLPLGGFTYGEKSLFLVNDWHAGLVPILLAAKYRPYGVYKDARSILIIHNLAHQGVEPAATYTNLGLPSEWYGA VGWVFPTWARTHALDTGEAVNVLKGAIVTSDRIITVSQGYAWEITTVEGGYGLQDLLSSRKSVINGITNGINVDEWNPSTDEHIPFHYSADDFSEKVKCK MALQKELGLPIRPECPMIGFIGRLDYQKGIDLIQTAGPDLMVDDIQFVMLGSGDPKYESWMRSMEETYRDKFRGWVGFNVPISHRITAGCDILLMPSRFE PCGLNQLYAMRYGTIPVVHGTGGLRDTVENFNPYAEGGAGAGTGWVFTPLSKDSMVSALRLAAATYREYKESWEGLMRRGMTRNYSWENAAVQYEQVFQ | |||
| Full Sequence |
|---|
| Protein Sequence Length: 656 Download |
| MASLQISGSV KFEPLVGFNR IRYFRPIGSL GIPRFRRRFS IGRPLLLRRS SSFSGDKIGE 60 SVGDEKGFIT DAERDGSGSV LGFQLTPTGD QEIFSTSTGE ITHQEEKKEA IDETVMADFG 120 VPGNRAVEEG AVEVGIPSGK AEVVNNLVFV TSEAAPYSKT GGLGDVCGSL PIALAGRGHR 180 VMVVSPRYLN GTAADNNYAR AKDLGIRITV NCFGGSQEVS FYHEYRDGVD WVFVDHKSYH 240 RPGNPYGDSK GAFGDNQFRF TLLCHAACEA PLVLPLGGFT YGEKSLFLVN DWHAGLVPIL 300 LAAKYRPYGV YKDARSILII HNLAHQGVEP AATYTNLGLP SEWYGAVGWV FPTWARTHAL 360 DTGEAVNVLK GAIVTSDRII TVSQGYAWEI TTVEGGYGLQ DLLSSRKSVI NGITNGINVD 420 EWNPSTDEHI PFHYSADDFS EKVKCKMALQ KELGLPIRPE CPMIGFIGRL DYQKGIDLIQ 480 TAGPDLMVDD IQFVMLGSGD PKYESWMRSM EETYRDKFRG WVGFNVPISH RITAGCDILL 540 MPSRFEPCGL NQLYAMRYGT IPVVHGTGGL RDTVENFNPY AEGGAGAGTG WVFTPLSKDS 600 MVSALRLAAA TYREYKESWE GLMRRGMTRN YSWENAAVQY EQVFQWVFMD PPYVS* 660 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK14099 | PRK14099 | 1.0e-105 | 150 | 645 | 506 | + glycogen synthase; Provisional | ||
| COG0297 | GlgA | 6.0e-145 | 146 | 647 | 509 | + Glycogen synthase [Carbohydrate transport and metabolism] | ||
| PRK00654 | glgA | 2.0e-173 | 146 | 645 | 514 | + glycogen synthase; Provisional | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 0 | 146 | 646 | 501 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| TIGR02095 | glgA | 0 | 146 | 645 | 501 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAF24126.1 | 0 | 88 | 655 | 8 | 575 | AF121673_1 soluble starch synthase [Arabidopsis thaliana] |
| GenBank | ABV25893.1 | 0 | 24 | 655 | 25 | 633 | starch synthase isoform I [Manihot esculenta] |
| RefSeq | NP_197818.1 | 0 | 1 | 655 | 1 | 652 | SSI1 (SUPPRESSOR OF SALICYLIC ACID INSENSITIVITY 1); starch synthase/ transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | XP_002277372.1 | 0 | 25 | 655 | 21 | 633 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002321996.1 | 0 | 1 | 655 | 1 | 649 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3vuf_A | 0 | 146 | 644 | 11 | 508 | A Chain A, Pectin Methylesterase Pema From Erwinia Chrysanthemi |
| PDB | 3vue_A | 0 | 146 | 644 | 11 | 508 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| PDB | 1rzv_B | 0 | 146 | 644 | 2 | 472 | A Chain A, Crystal Structure Of The Glycogen Synthase From Agrobacterium Tumefaciens (Non-Complexed Form) |
| PDB | 1rzv_A | 0 | 146 | 644 | 2 | 472 | A Chain A, Crystal Structure Of The Glycogen Synthase From Agrobacterium Tumefaciens (Non-Complexed Form) |
| PDB | 1rzu_B | 0 | 146 | 644 | 2 | 472 | A Chain A, Crystal Structure Of The Glycogen Synthase From A. Tumefaciens In Complex With Adp |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO595058 | 492 | 162 | 653 | 0 |
| HO581986 | 368 | 255 | 621 | 0 |
| GO872237 | 347 | 306 | 652 | 0 |
| CK120636 | 260 | 68 | 327 | 0 |
| HO581986 | 34 | 620 | 653 | 0.00002 |
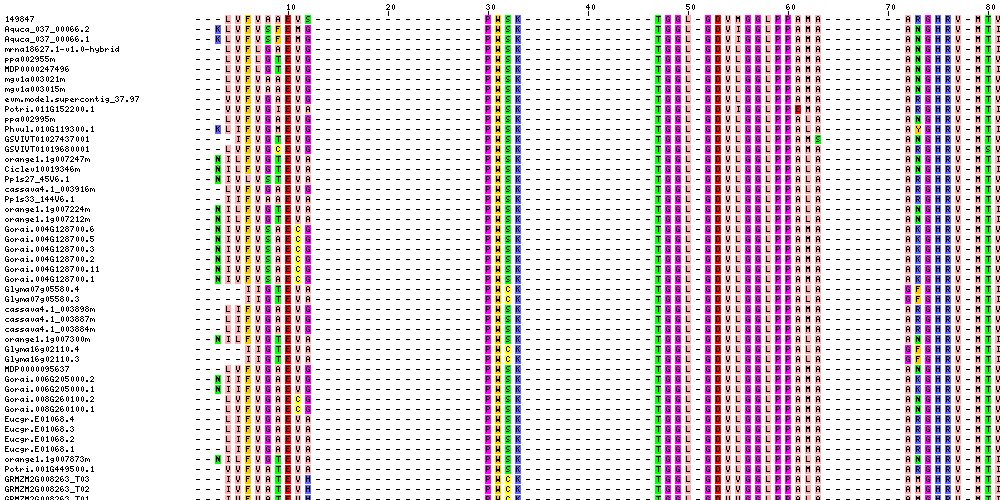
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |