

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s2_195V6.1 |
| Family | GT5 |
| Protein Properties | Length: 887 Molecular Weight: 99811.9 Isoelectric Point: 4.9491 |
| Chromosome | Chromosome/Scaffold: 2 Start: 979022 End: 986610 |
| Description | starch synthase 4 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 393 | 879 | 0 |
| VIHIAAEMAPVAKVGGLGDVLTGLSRSLQRKGHLVEAILPKYDCMDYSRISNLKELDLELHSSFDGQTHKNKVWSGIVEGLPVYFIEPLHPAKFFWRGQF YSEPDDFRRFTYFCRAAMEFLLQSGKRPDIIHSHDWQTAVVAPLYWDVFVPLGLDSARLAFTCHNFEYQGAENSGAIAACGLSPQTLMRPDRMQDNSHHN KINLLKGGVVFSNVVTTVSPTYAQEVLRPEGGKGLHGTLGVHSKKFFGVLNGIDQEVWDPATDTLIDCQYSAHDIDGKFVNKQRLRERLGLASEGEDERR PLVGCVTRLVPQKGVHLIRHAIYRTLEKGGQFILLGSSPVPHIQHEFEGIARQFENHPQIRLVLKYDEALSHAIYAASDIFVIPSIFEPCGLTQMISMRY GTIPVVRRTGGLADSVFDVDDERNPEEKKNGFVFDDANEGSLNWGLDRALDYYSQRPEWWQDLVKKAMLIDFSWDTSADQYVDLYKQ | |||
| Full Sequence |
|---|
| Protein Sequence Length: 887 Download |
| MPASTANRES TGVSVNDLML LVKEAETNIR TLSQLRERAV NELQTARNEK EELQSQVQIL 60 KARLAESEAK LKYTVQARER TQLLEEEVEA LRQGLAQTEV DSQAFATLKE ENRVMKESLT 120 ALKQAGGNGL LEAKYQSSLD EVNSLRVQLA DAQAEITKFS RGTGRERAGL QEYIVGLEKK 180 IADMEKQLAE ANANSAELKS LEDESMALQE EAAALRAQLA GRSDSSSSSE DQTLQRQVDS 240 LKSLLARAVA EKNALLDLRS ENLRLREQVK LLEERIRESD AEIQAQLQIY TAEVEAFQAS 300 LNDLKSENET SVVQVPVDEM TWDFWSELLL RIDALMLGDL LTQEDAKELH LMAWRREKRL 360 CDVYVNLADE PDEDLVGAFQ ELLQPKKRPG IYVIHIAAEM APVAKVGGLG DVLTGLSRSL 420 QRKGHLVEAI LPKYDCMDYS RISNLKELDL ELHSSFDGQT HKNKVWSGIV EGLPVYFIEP 480 LHPAKFFWRG QFYSEPDDFR RFTYFCRAAM EFLLQSGKRP DIIHSHDWQT AVVAPLYWDV 540 FVPLGLDSAR LAFTCHNFEY QGAENSGAIA ACGLSPQTLM RPDRMQDNSH HNKINLLKGG 600 VVFSNVVTTV SPTYAQEVLR PEGGKGLHGT LGVHSKKFFG VLNGIDQEVW DPATDTLIDC 660 QYSAHDIDGK FVNKQRLRER LGLASEGEDE RRPLVGCVTR LVPQKGVHLI RHAIYRTLEK 720 GGQFILLGSS PVPHIQHEFE GIARQFENHP QIRLVLKYDE ALSHAIYAAS DIFVIPSIFE 780 PCGLTQMISM RYGTIPVVRR TGGLADSVFD VDDERNPEEK KNGFVFDDAN EGSLNWGLDR 840 ALDYYSQRPE WWQDLVKKAM LIDFSWDTSA DQYVDLYKQV LSKANA* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02316 | PLN02316 | 2.0e-153 | 387 | 877 | 498 | + synthase/transferase | ||
| PLN02939 | PLN02939 | 0 | 17 | 886 | 882 | + transferase, transferring glycosyl groups | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 0 | 393 | 879 | 492 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| TIGR02095 | glgA | 0 | 391 | 880 | 493 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| PRK00654 | glgA | 0 | 392 | 883 | 498 | + glycogen synthase; Provisional | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAB40375.1 | 0 | 283 | 886 | 269 | 870 | starch synthase, isoform V [Vigna unguiculata] |
| RefSeq | NP_193558.3 | 0 | 161 | 886 | 307 | 1035 | ATSS4; transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | XP_001751723.1 | 0 | 71 | 886 | 44 | 835 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001756311.1 | 0 | 283 | 886 | 26 | 667 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001759580.1 | 0 | 237 | 881 | 1 | 643 | predicted protein [Physcomitrella patens subsp. patens] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3d1j_A | 0 | 393 | 880 | 3 | 476 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 3guh_A | 0 | 393 | 880 | 3 | 476 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2r4u_A | 0 | 393 | 880 | 3 | 476 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2r4t_A | 0 | 393 | 880 | 3 | 476 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 2qzs_A | 0 | 393 | 880 | 3 | 476 | A Chain A, Crystal Structure Of Wild-Type E.Coli Gs In Complex With Adp And Glucose(Wtgsb) |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| starch biosynthesis | GLYCOGENSYN-RXN | EC-2.4.1.21 | starch synthase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| GO851132 | 335 | 466 | 800 | 0 |
| FC341609 | 271 | 114 | 384 | 0 |
| CF828776 | 288 | 516 | 803 | 0 |
| GW865616 | 300 | 497 | 796 | 0 |
| CB893818 | 281 | 390 | 670 | 0 |
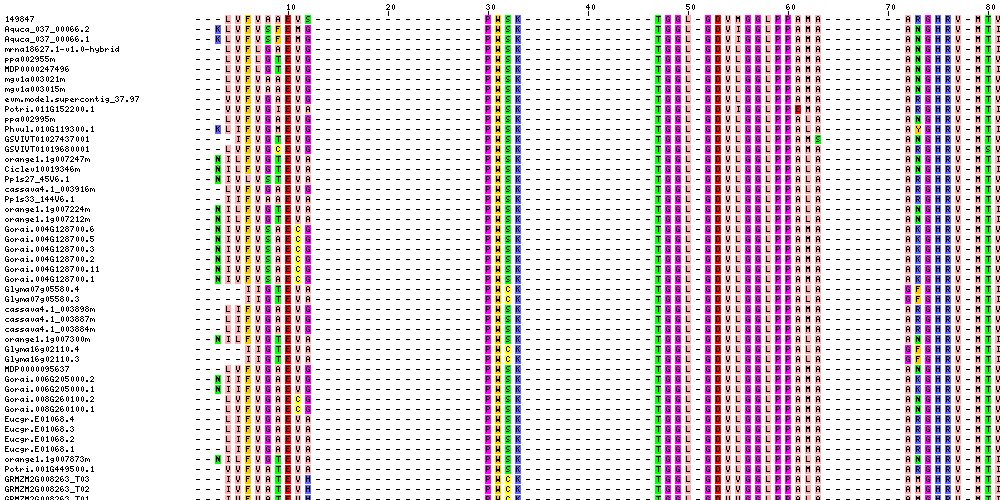
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |