

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s34_269V6.1 |
| Family | GT5 |
| Protein Properties | Length: 812 Molecular Weight: 91658.2 Isoelectric Point: 6.8566 |
| Chromosome | Chromosome/Scaffold: 34 Start: 1374871 End: 1382391 |
| Description | starch synthase 4 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 324 | 805 | 0 |
| HIVHICTEMAPVAQPGTASFNISNLCKALRRNGHLVEVILPKYDCMDLSAVDNLREIETDLYIYFGDQWHKNRIWIGTLYGIAVTLLEPFHPGNFFAREE FYNYNDDFERFTYFCKLSLEYLLKLGKQADILHLHNWQTAAVAPMFWEHYAHQGLQDIRLVLTCHDFRYQCLRDPHKLAICGLDPEQLNRPDRLQDDLEP RFINMLKGGIVYSNIVTTVSPIYAADVLTKEHGYDLHVTLNTHKDKFIGIYNGLDDVLWDPALDCALPETYSVEDMAGKSVCKASLRQELGLPMVDDSVP LVGCILFENSEVELKLIRAALECAVGEEAQFVCAVHNNPGLGIRLTGKQIHTDANARLLGVNDDALLHRIVAASDILYSPDMYEPASQSVMIGMRYGAVP VARQVHSITSSIINLDEQQHIILHATGFPFYSTEDSDAITSLHDALTYWKCHPEMWSTLVRNCMSKDLSWDSSCVEAYEAAY | |||
| Full Sequence |
|---|
| Protein Sequence Length: 812 Download |
| MSVMQHMAAA NNKWHAAALA LPAAPRGFGR RHRVSCCIQP PVGDRGIGED RLTEISIATD 60 GDGIFGVDMK QLIDDAQQSK CMADGELKSC QIATIFHLLH NTLFTRSQKF SYIRALIIVC 120 NNVEDRLEAD KRELSRFSLK NSRVHDLHHK VSQLTCRNIP GVKGMPFKMP GILHLNTGRT 180 TAFEKLQKVE RERDLLAARV AQLKAEHEAS MAECDLLRQK AKQFEMSFHV SGRQGKSAGS 240 MKRFKMNRKR FNPPSIWSEI LLRIDAMTLT GVLQIEQATS LRKLVWARDA QIADMFFKLA 300 DGNDAELSAR LLSIISSTRR RSLHIVHICT EMAPVAQPGT ASFNISNLCK ALRRNGHLVE 360 VILPKYDCMD LSAVDNLREI ETDLYIYFGD QWHKNRIWIG TLYGIAVTLL EPFHPGNFFA 420 REEFYNYNDD FERFTYFCKL SLEYLLKLGK QADILHLHNW QTAAVAPMFW EHYAHQGLQD 480 IRLVLTCHDF RYQCLRDPHK LAICGLDPEQ LNRPDRLQDD LEPRFINMLK GGIVYSNIVT 540 TVSPIYAADV LTKEHGYDLH VTLNTHKDKF IGIYNGLDDV LWDPALDCAL PETYSVEDMA 600 GKSVCKASLR QELGLPMVDD SVPLVGCILF ENSEVELKLI RAALECAVGE EAQFVCAVHN 660 NPGLGIRLTG KQIHTDANAR LLGVNDDALL HRIVAASDIL YSPDMYEPAS QSVMIGMRYG 720 AVPVARQVHS ITSSIINLDE QQHIILHATG FPFYSTEDSD AITSLHDALT YWKCHPEMWS 780 TLVRNCMSKD LSWDSSCVEA YEAAYWSVHK V* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0297 | GlgA | 1.0e-77 | 325 | 805 | 490 | + Glycogen synthase [Carbohydrate transport and metabolism] | ||
| PRK00654 | glgA | 3.0e-100 | 324 | 805 | 505 | + glycogen synthase; Provisional | ||
| TIGR02095 | glgA | 9.0e-110 | 324 | 805 | 488 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 9.0e-110 | 324 | 807 | 503 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| PLN02939 | PLN02939 | 1.0e-156 | 194 | 805 | 621 | + transferase, transferring glycosyl groups | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACD13789.1 | 0 | 255 | 805 | 131 | 676 | starch synthase V precursor [Vitis vinifera] |
| RefSeq | XP_001751723.1 | 0 | 256 | 797 | 273 | 819 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001758914.1 | 0 | 171 | 811 | 11 | 651 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_002274716.1 | 0 | 253 | 805 | 444 | 998 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002310316.1 | 0 | 252 | 805 | 117 | 663 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3guh_A | 2e-36 | 323 | 808 | 1 | 476 | A Chain A, Crystal Strucuture Of An Inositol Monophosphatase Family Protein (Sas2203) From Staphylococcus Aureus Mssa476 |
| PDB | 2r4u_A | 2e-36 | 323 | 808 | 1 | 476 | A Chain A, Crystal Strucuture Of An Inositol Monophosphatase Family Protein (Sas2203) From Staphylococcus Aureus Mssa476 |
| PDB | 2r4t_A | 2e-36 | 323 | 808 | 1 | 476 | A Chain A, Crystal Strucuture Of An Inositol Monophosphatase Family Protein (Sas2203) From Staphylococcus Aureus Mssa476 |
| PDB | 2qzs_A | 2e-36 | 323 | 808 | 1 | 476 | A Chain A, Crystal Structure Of Wild-Type E.Coli Gs In Complex With Adp And Glucose(Wtgsb) |
| PDB | 3cx4_A | 2e-35 | 323 | 806 | 1 | 474 | A Chain A, Crystal Structure Of Wild-Type E.Coli Gs In Complex With Adp And Glucose(Wtgsb) |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| starch biosynthesis | GLYCOGENSYN-RXN | EC-2.4.1.21 | starch synthase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| FC360455 | 269 | 471 | 739 | 0 |
| FC328247 | 255 | 178 | 432 | 0 |
| FC360377 | 251 | 201 | 451 | 0 |
| FC358107 | 245 | 260 | 504 | 0 |
| FC371371 | 255 | 171 | 424 | 0 |
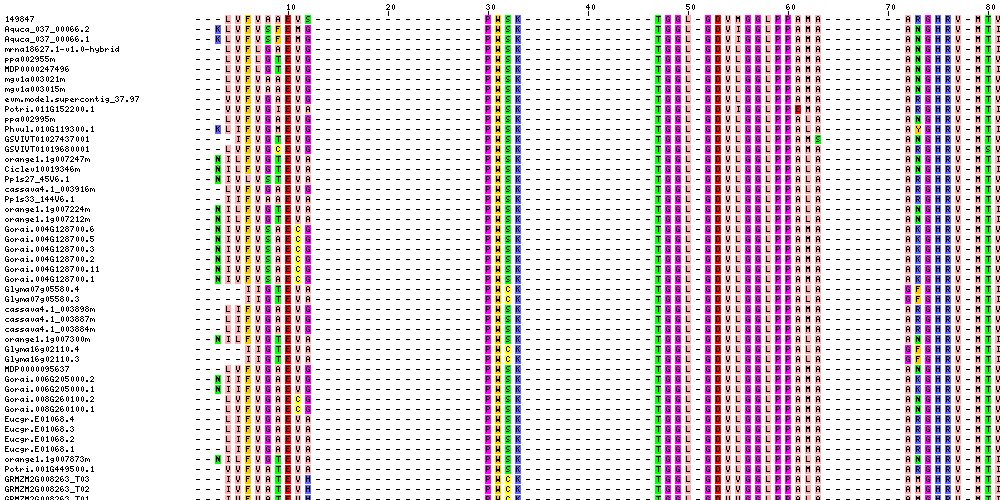
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |