

| Basic Information | |
|---|---|
| Species | Citrus sinensis |
| Cazyme ID | orange1.1g008028m |
| Family | GT5 |
| Protein Properties | Length: 581 Molecular Weight: 63954.9 Isoelectric Point: 9.0777 |
| Chromosome | Chromosome/Scaffold: 00009 Start: 2211506 End: 2214354 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 87 | 551 | 0 |
| LVFVGAEVAPWSKTGGLGDVLGGLPPALAARGHRVMSVAPRYDQYKDAWDTSVLAEVKVGDSIETVRFFHCYKRGVDRVFVDHPVFLEKVWGKTGSKIYG PKAGLDYEDNQLRFSLLCQAALEAPRILNLNNNKYFSGPYGEDVLFIGNDWHTALLPCYLKFMYQSKGIHKNAKRLCRPRYDKPVKGRKINWMKAGILES DRVLTVSPHYAKELISGEEKGVELDNIIRKTGISGIVNGMDVQEWNPSTDRYINVNYDATTVMNAKPLVKEALQSQLGLPVDRNIPLIGFIGRLEEQKGS DILAQAIPKFMGGNVQIVVLGTGKKPMEQQIEQLEIICPEKARGITKFSTPLAHKIIAGADFMLVPSRFEPCGLIQLHAMRYGTVPIVASTGGLFDTVKE GITGFQMRSFHVECDRVDLADVAAIAKHVNRAVATYGTPALKEMIQNCMALDLSWKEPARLWEKM | |||
| Full Sequence |
|---|
| Protein Sequence Length: 581 Download |
| MATMTAPQFI GGSSQLTSGS VIKSNFSQIG LRSQSMTHSG LRSLNTIDRL QMKSYAKAVV 60 TKATKNEQQT PNNVPSKKII CQQGMNLVFV GAEVAPWSKT GGLGDVLGGL PPALAARGHR 120 VMSVAPRYDQ YKDAWDTSVL AEVKVGDSIE TVRFFHCYKR GVDRVFVDHP VFLEKVWGKT 180 GSKIYGPKAG LDYEDNQLRF SLLCQAALEA PRILNLNNNK YFSGPYGEDV LFIGNDWHTA 240 LLPCYLKFMY QSKGIHKNAK RLCRPRYDKP VKGRKINWMK AGILESDRVL TVSPHYAKEL 300 ISGEEKGVEL DNIIRKTGIS GIVNGMDVQE WNPSTDRYIN VNYDATTVMN AKPLVKEALQ 360 SQLGLPVDRN IPLIGFIGRL EEQKGSDILA QAIPKFMGGN VQIVVLGTGK KPMEQQIEQL 420 EIICPEKARG ITKFSTPLAH KIIAGADFML VPSRFEPCGL IQLHAMRYGT VPIVASTGGL 480 FDTVKEGITG FQMRSFHVEC DRVDLADVAA IAKHVNRAVA TYGTPALKEM IQNCMALDLS 540 WKEPARLWEK MLLSMEVAGS EPGIEGDEIA PLARENLATP * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PRK14099 | PRK14099 | 7.0e-72 | 181 | 551 | 402 | + glycogen synthase; Provisional | ||
| COG0297 | GlgA | 7.0e-91 | 85 | 552 | 511 | + Glycogen synthase [Carbohydrate transport and metabolism] | ||
| PRK00654 | glgA | 2.0e-112 | 85 | 551 | 509 | + glycogen synthase; Provisional | ||
| TIGR02095 | glgA | 2.0e-145 | 85 | 554 | 505 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 3.0e-155 | 86 | 552 | 505 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACH72975.1 | 0 | 1 | 580 | 1 | 615 | granule-bound starch synthase [Nelumbo nucifera] |
| GenBank | ACJ11735.1 | 0 | 1 | 580 | 1 | 609 | granule bound starch synthase [Gossypium hirsutum] |
| GenBank | ACJ11751.1 | 0 | 1 | 580 | 1 | 609 | granule bound starch synthase [Gossypium hirsutum] |
| GenBank | ACM78591.1 | 0 | 1 | 580 | 1 | 615 | granule-bound starch synthase [Nelumbo nucifera] |
| GenBank | ACV72639.1 | 0 | 1 | 580 | 1 | 609 | granule-bound starch synthase 1 [Gossypium hirsutum] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3vuf_A | 0 | 85 | 580 | 10 | 536 | B Chain B, Crystal Structure Analysis Of Coagulation Factor Viii |
| PDB | 3vue_A | 0 | 85 | 580 | 10 | 536 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| PDB | 3guh_A | 0 | 85 | 557 | 1 | 478 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| PDB | 2r4u_A | 0 | 85 | 557 | 1 | 478 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| PDB | 2r4t_A | 0 | 85 | 557 | 1 | 478 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| AJ568414 | 616 | 1 | 581 | 0 |
| HO629148 | 586 | 30 | 581 | 0 |
| HO825853 | 518 | 93 | 579 | 0 |
| ES793310 | 315 | 267 | 581 | 0 |
| HO778340 | 394 | 61 | 421 | 0 |
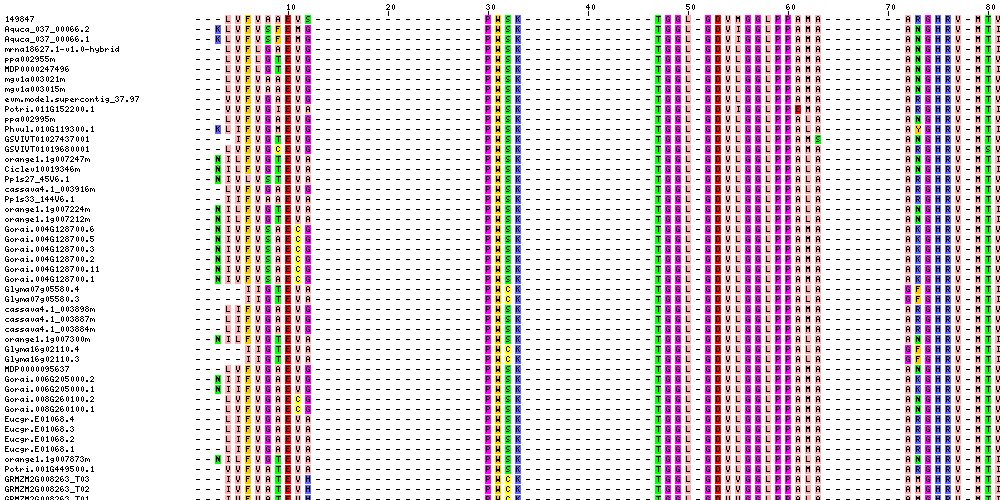
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |