

| Basic Information | |
|---|---|
| Species | Citrus sinensis |
| Cazyme ID | orange1.1g002589m |
| Family | GT5 |
| Protein Properties | Length: 905 Molecular Weight: 102897 Isoelectric Point: 6.2919 |
| Chromosome | Chromosome/Scaffold: 00007 Start: 1381600 End: 1386488 |
| Description | starch synthase 4 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 513 | 854 | 0 |
| HVIHIAAEMAPVAKVGGLGDVVAGLGKALQKKGHLVEIVLPKYDCMQYDRIDDLRALDVVVESYFDGRLFKNKVWVSTIEGLPVYFIEPHHPDKFFWRGQ FYGEHDDFRRFSFFSRAALELLLQAGKQPDIIHCHDWQTAFVAPLYWDLYVPKGLNSARVCFTCHNFEYQGTAPAKELASCGLDVQQLNRPDRMQDNSAH DRINPLKGAIVFSNIVTTVSPSYAQEVRTSEGGQGLHSTLNFHSKKFVGILNGIDTDAWNPATDTFLKVQYNANDLQGKAENKESIRKHLGLSSADARKP LVGCITRLVPQKGVHLIRHAIYRTLELGGQFILLGSSPVPHI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 905 Download |
| MASKISTSFI SPFVIHFNCK NSNNKNKHLN VPLLFSSRRL LPASCKMRQR SFGSQQKRQH 60 VKKGSPDQQR PNDADLVPTS DGDSESESSL IDREPIDVEH TEEQNLGSVF VPELKESLVL 120 NCDGGEELST SQLDNLISMI RNAEKNILLL NEARVQALED LHKILQEKEA LQGEINALEM 180 RLAETDARIR VAAQEKIHVE LLEDQLQKLQ HELTHRGVSE HSELDVFANQ NEPANEDLVL 240 NNSEIHSFSK ELDSLKTENL SLKNDIKVLK AELNSVKDAD ERVVMLEMER SSLESSLKEL 300 ESKLSISQED VAKLSTLKVE CKDLYEKVEN LQGLLAKATK QADQAISVLQ QNQELRKKVD 360 KLEESLDEAN IYKLSSEKMQ QYNELMQQKM KLLEERLQRS DEEIHSYVQL YQESVKEFQD 420 TLHSLKEESK KRAVHEPVDD MPWEFWSRLL LIIDGWLLEK KLSTSEAKLL REMVWKRNGR 480 IRDAYMECKE KNEHEAISTF LKLTSSSISS GLHVIHIAAE MAPVAKVGGL GDVVAGLGKA 540 LQKKGHLVEI VLPKYDCMQY DRIDDLRALD VVVESYFDGR LFKNKVWVST IEGLPVYFIE 600 PHHPDKFFWR GQFYGEHDDF RRFSFFSRAA LELLLQAGKQ PDIIHCHDWQ TAFVAPLYWD 660 LYVPKGLNSA RVCFTCHNFE YQGTAPAKEL ASCGLDVQQL NRPDRMQDNS AHDRINPLKG 720 AIVFSNIVTT VSPSYAQEVR TSEGGQGLHS TLNFHSKKFV GILNGIDTDA WNPATDTFLK 780 VQYNANDLQG KAENKESIRK HLGLSSADAR KPLVGCITRL VPQKGVHLIR HAIYRTLELG 840 GQFILLGSSP VPHIQVYPIL LSSFSFLRKH IFNICNLYIK LGQGGDLTVN NNCEPWLHHI 900 EVWC* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| PLN02316 | PLN02316 | 2.0e-80 | 512 | 855 | 345 | + synthase/transferase | ||
| PRK00654 | glgA | 2.0e-108 | 512 | 848 | 339 | + glycogen synthase; Provisional | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 2.0e-114 | 513 | 848 | 338 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| TIGR02095 | glgA | 2.0e-116 | 512 | 848 | 342 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| PLN02939 | PLN02939 | 0 | 30 | 856 | 842 | + transferase, transferring glycosyl groups | ||
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACT09058.1 | 0 | 37 | 855 | 33 | 847 | starch synthase IV precursor [Solanum lycopersicum] |
| RefSeq | NP_193558.3 | 6e-18 | 1 | 167 | 1 | 219 | ATSS4; transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | NP_193558.3 | 0 | 273 | 855 | 303 | 885 | ATSS4; transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | XP_002274716.1 | 0 | 1 | 856 | 1 | 858 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002298514.1 | 0 | 102 | 855 | 1 | 733 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1rzv_B | 4e-38 | 512 | 851 | 1 | 331 | A Chain A, Crystal Structure Of The Glycogen Synthase From Agrobacterium Tumefaciens (Non-Complexed Form) |
| PDB | 1rzv_A | 4e-38 | 512 | 851 | 1 | 331 | A Chain A, Crystal Structure Of The Glycogen Synthase From Agrobacterium Tumefaciens (Non-Complexed Form) |
| PDB | 1rzu_B | 4e-38 | 512 | 851 | 1 | 331 | A Chain A, Crystal Structure Of The Glycogen Synthase From A. Tumefaciens In Complex With Adp |
| PDB | 1rzu_A | 4e-38 | 512 | 851 | 1 | 331 | A Chain A, Crystal Structure Of The Glycogen Synthase From A. Tumefaciens In Complex With Adp |
| PDB | 3d1j_A | 3e-35 | 512 | 851 | 1 | 333 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DV124304 | 258 | 576 | 833 | 0 |
| EL448877 | 316 | 440 | 755 | 0 |
| CB893818 | 284 | 508 | 791 | 0 |
| EG664148 | 262 | 542 | 803 | 0 |
| CF828776 | 219 | 637 | 855 | 0 |
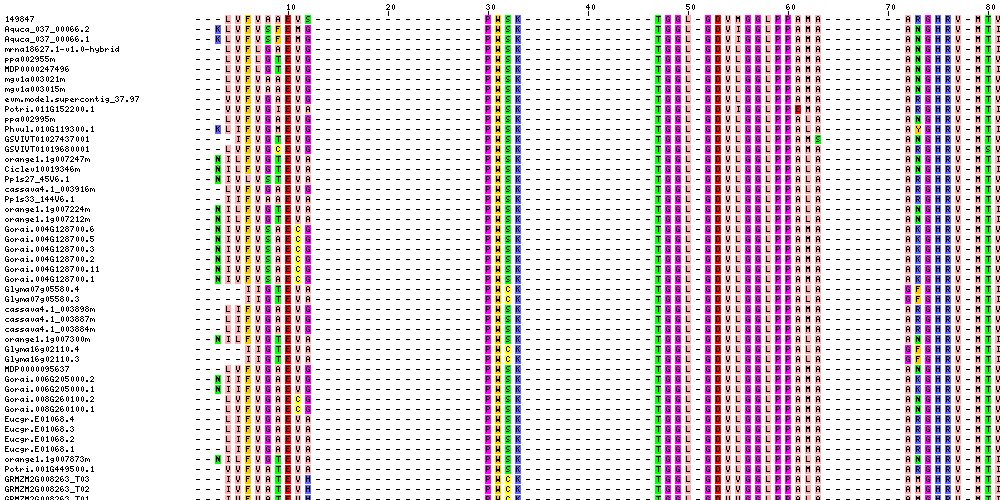
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |