

| Basic Information | |
|---|---|
| Species | Gossypium raimondii |
| Cazyme ID | Gorai.004G128700.4 |
| Family | GT5 |
| Protein Properties | Length: 551 Molecular Weight: 60721.9 Isoelectric Point: 8.9669 |
| Chromosome | Chromosome/Scaffold: 04 Start: 34283494 End: 34287287 |
| Description | UDP-Glycosyltransferase superfamily protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 84 | 380 | 0 |
| NIVFVSAECGPWSKTGGLGDVLGGLPPAMAAKGHRVMTVCPRYDQYKDAWDTSVLVDLKVGDKVVTVRFFHCYKRGVDRVFVDHPMFLEKVWGKTASKIY GPRAGLDYEDNQLRFSLLCQAALEAPRVLNLNSSKNFSGPYGEDVVFIANDWHSALLPCYLKSMYQSRGIYMNAKVVFCIHNIAYQGRFAFADFKRLNLP ERFKSSFDFIDGYNKPVKGRKINWMKAGILESHRVLTVSPYYAQELVSGEDKGVELDNIIRKTGITGIVNGMDVQEWNPASDKYISVKYDATTGTGK | |||
| GT5 | 376 | 521 | 0 |
| TGTGKKAMEKQIEQLEIQYPDNVRAVAKFNVSLAHMIIAGADYILVPSRFEPCGLIQLHAMRYGTVPIVASTGGLVDTVKEGFTGFQMGAFNVECDEVDP SDVIKVVKTVKRALATYGTQALKEMIQNCMAQDFSWKGPSRLWEKM | |||
| Full Sequence |
|---|
| Protein Sequence Length: 551 Download |
| MATLTTSHFV STSSHFSSHG ADTKANLAQV GARNQAMTHN GLRSLNKVDR LQIRTTNAKA 60 VVTKAMKQAD HRPLGKIICG IGMNIVFVSA ECGPWSKTGG LGDVLGGLPP AMAAKGHRVM 120 TVCPRYDQYK DAWDTSVLVD LKVGDKVVTV RFFHCYKRGV DRVFVDHPMF LEKVWGKTAS 180 KIYGPRAGLD YEDNQLRFSL LCQAALEAPR VLNLNSSKNF SGPYGEDVVF IANDWHSALL 240 PCYLKSMYQS RGIYMNAKVV FCIHNIAYQG RFAFADFKRL NLPERFKSSF DFIDGYNKPV 300 KGRKINWMKA GILESHRVLT VSPYYAQELV SGEDKGVELD NIIRKTGITG IVNGMDVQEW 360 NPASDKYISV KYDATTGTGK KAMEKQIEQL EIQYPDNVRA VAKFNVSLAH MIIAGADYIL 420 VPSRFEPCGL IQLHAMRYGT VPIVASTGGL VDTVKEGFTG FQMGAFNVEC DEVDPSDVIK 480 VVKTVKRALA TYGTQALKEM IQNCMAQDFS WKGPSRLWEK MLLSLGVAGS EPGIEGEEVA 540 PLAKENVATP * |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 4.0e-48 | 377 | 522 | 153 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| TIGR02095 | glgA | 4.0e-50 | 377 | 525 | 151 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| TIGR02095 | glgA | 5.0e-79 | 83 | 398 | 323 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| cd03791 | GT1_Glycogen_synthase_DULL1_like | 1.0e-81 | 84 | 382 | 305 | + This family is most closely related to the GT1 family of glycosyltransferases. Glycogen synthase catalyzes the formation and elongation of the alpha-1,4-glucose backbone using ADP-glucose, the second and key step of glycogen biosynthesis. This family includes starch synthases of plants, such as DULL1 in Zea mays and glycogen synthases of various organisms. | ||
| PRK00654 | glgA | 7.0e-96 | 83 | 511 | 497 | + glycogen synthase; Provisional | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ACJ11735.1 | 0 | 1 | 550 | 1 | 609 | granule bound starch synthase [Gossypium hirsutum] |
| GenBank | ACJ11751.1 | 0 | 1 | 550 | 1 | 609 | granule bound starch synthase [Gossypium hirsutum] |
| GenBank | ACV72639.1 | 0 | 1 | 550 | 1 | 609 | granule-bound starch synthase 1 [Gossypium hirsutum] |
| Swiss-Prot | O82627 | 0 | 1 | 550 | 1 | 608 | SSG1_ANTMA RecName: Full=Granule-bound starch synthase 1, chloroplastic/amyloplastic; AltName: Full=Granule-bound starch synthase I; Short=GBSS-I; Flags: Precursor |
| Swiss-Prot | Q43784 | 0 | 1 | 550 | 1 | 608 | SSG1_MANES RecName: Full=Granule-bound starch synthase 1, chloroplastic/amyloplastic; AltName: Full=Granule-bound starch synthase I; Short=GBSS-I; Flags: Precursor |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3vuf_A | 0 | 83 | 395 | 10 | 322 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 3vuf_A | 0 | 377 | 550 | 363 | 536 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| PDB | 3vue_A | 0 | 83 | 395 | 10 | 322 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| PDB | 3vue_A | 0 | 377 | 550 | 363 | 536 | A Chain A, Crystal Structure Of Rice Granule Bound Starch Synthase I Catalytic Domain |
| PDB | 3d1j_A | 3e-32 | 83 | 525 | 1 | 476 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| AJ568414 | 610 | 1 | 551 | 0 |
| HO825853 | 518 | 91 | 549 | 0 |
| ES813520 | 330 | 24 | 350 | 0 |
| CO114733 | 266 | 219 | 484 | 0 |
| CO125235 | 284 | 59 | 342 | 0 |
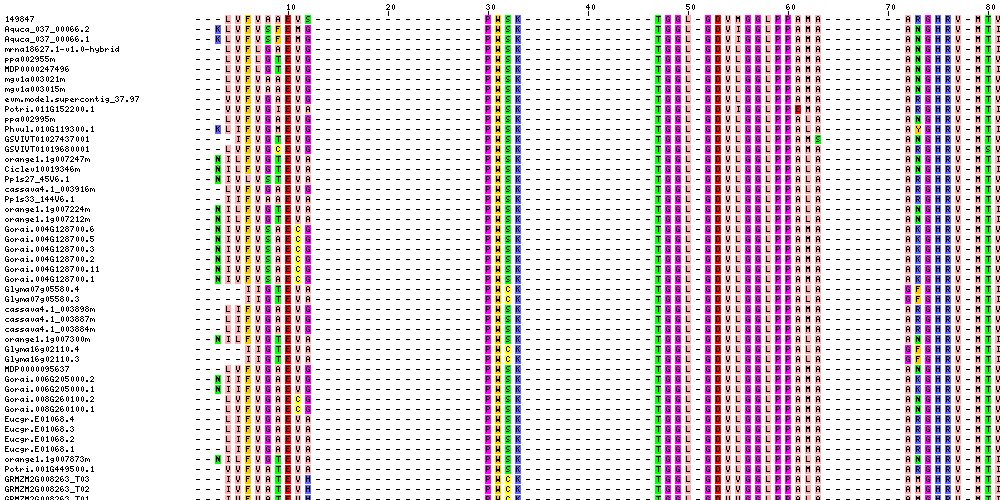
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |