

| Basic Information | |
|---|---|
| Species | Aquilegia coerulea |
| Cazyme ID | Aquca_007_00320.4 |
| Family | GH35 |
| Protein Properties | Length: 696 Molecular Weight: 77799.1 Isoelectric Point: 6.8649 |
| Chromosome | Chromosome/Scaffold: 7 Start: 2860282 End: 2866090 |
| Description | beta-galactosidase 3 |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH35 | 38 | 340 | 0 |
| IINGQRRILISGSIHYPRSTPDMWEDLIMKAKEGGLDVIQTYVFWNGHEPSPGNYYFEGRYDLVRFIKTVQKAGLYVHLRIGPYVCAEWNFGGFPVWLKY VPGISFRTDNEPFKMAMQGFTQKIVQMMKSESLYETQGGPIILSQIENEYGSEAKAFGAAGQAYMNWAATMAVRLDTGVPWVMCKEDDAPDPVINACNGF YCDAFTPNKPYKPTMWTEAWSGWFTEFGGTVHQRPVEDLAFAVARFIQKGGSFINYYMYHGGTNFGRTAGGPFITTSYDYDAPIDEYGLTRQPKYGHLKE LHR | |||
| Full Sequence |
|---|
| Protein Sequence Length: 696 Download |
| METNSISKSI FFLYFCFIIL VNLQLIQCGV TYDKKAIIIN GQRRILISGS IHYPRSTPDM 60 WEDLIMKAKE GGLDVIQTYV FWNGHEPSPG NYYFEGRYDL VRFIKTVQKA GLYVHLRIGP 120 YVCAEWNFGG FPVWLKYVPG ISFRTDNEPF KMAMQGFTQK IVQMMKSESL YETQGGPIIL 180 SQIENEYGSE AKAFGAAGQA YMNWAATMAV RLDTGVPWVM CKEDDAPDPV INACNGFYCD 240 AFTPNKPYKP TMWTEAWSGW FTEFGGTVHQ RPVEDLAFAV ARFIQKGGSF INYYMYHGGT 300 NFGRTAGGPF ITTSYDYDAP IDEYGLTRQP KYGHLKELHR AIKLSERALV SADPIVTSLG 360 TYQQAHVFTS GSGDCAAFLS NFNSKSAARV MFNNMHYNLP PWSISILPDC RNAVFNTAKV 420 GVQTSQMQML PSNSQLLSWE TFDEDMSSLG EDRMINSFGL LEQINVTRDS SDYLWYITSV 480 EINPSESFLN GGEWPTLTVG SKGHAMHVFI NGQLSGSARG TRENRRFTYT GKVNMRAGTN 540 RIALLSIAVG LPNNGAHYET WSTGVLGPVE LHGLKEGKRD LTWQKWSYQV GLKGEAMNVV 600 SPNGVSSAEW MQASLAAQKQ QPLTWFKTHF DEPSGNEPLA LDMGSMGKGQ VWVNGESIGR 660 YWTAYAEGNC NECSYSGTYR PPKCQVGCGQ PTQRW* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
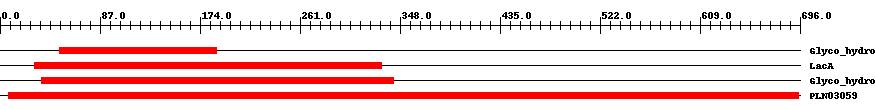 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam02449 | Glyco_hydro_42 | 2.0e-5 | 52 | 188 | 153 | + Beta-galactosidase. This group of beta-galactosidase enzymes belong to the glycosyl hydrolase 42 family. The enzyme catalyzes the hydrolysis of terminal, non-reducing terminal beta-D-galactosidase residues. | ||
| COG1874 | LacA | 6.0e-20 | 30 | 332 | 351 | + Beta-galactosidase [Carbohydrate transport and metabolism] | ||
| pfam01301 | Glyco_hydro_35 | 2.0e-168 | 36 | 342 | 322 | + Glycosyl hydrolases family 35. | ||
| PLN03059 | PLN03059 | 0 | 7 | 695 | 692 | + beta-galactosidase; Provisional | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0004565 | beta-galactosidase activity |
| GO:0005975 | carbohydrate metabolic process |
| GO:0009341 | beta-galactosidase complex |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAK81874.1 | 0 | 1 | 695 | 1 | 694 | putative beta-galactosidase BG1 [Vitis vinifera] |
| DDBJ | BAF31234.1 | 0 | 1 | 695 | 3 | 695 | beta-D-galactosidase [Persea americana] |
| EMBL | CAN78072.1 | 0 | 1 | 695 | 1 | 694 | hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002274449.1 | 0 | 1 | 695 | 1 | 693 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002280715.1 | 0 | 1 | 695 | 1 | 694 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3d3a_A | 0 | 34 | 338 | 12 | 325 | A Chain A, Crystal Structure Of A Beta-Galactosidase From Bacteroides Thetaiotaomicron |
| PDB | 3thd_D | 3.99931e-42 | 30 | 354 | 11 | 347 | A Chain A, Crystal Structure Of A Beta-Galactosidase From Bacteroides Thetaiotaomicron |
| PDB | 3thd_C | 3.99931e-42 | 30 | 354 | 11 | 347 | A Chain A, Crystal Structure Of A Beta-Galactosidase From Bacteroides Thetaiotaomicron |