

| Basic Information | |
|---|---|
| Species | Medicago truncatula |
| Cazyme ID | Medtr7g088740.1 |
| Family | CBM20 |
| Protein Properties | Length: 1023 Molecular Weight: 117660 Isoelectric Point: 5.9091 |
| Chromosome | Chromosome/Scaffold: 7 Start: 27411865 End: 27421513 |
| Description | disproportionating enzyme 2 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CBM20 | 16 | 104 | 3.2e-23 |
| VKISFRLPYLTQWGQSLLVCGSVPVLGSWNVKKGVLLSPFHEGSELIWSGSITVPKGFQCEYTYYVVDDKKNIVRWEMGKKHELALPDG | |||
| GH77 | 813 | 966 | 1.4e-35 |
| CVMQELGLVGLRIQRMPNESDLEFGIPSQYSYMTVCAPSCHDCSTLRAWWEEDEERRQRFFKNVMESDELPPDQCVPEVAHFIIRQHIESPSMWAIFPLQ DLLALKEEYTARPAIEETINDPTNPKHYWRYRVHVTLESLNKDNELKTIIKDLV | |||
| GH77 | 278 | 801 | 0 |
| IPMFSIRSESDLGVGEFLDLKLLVDWAVASGFHLVQLLPINDTSVHQMWWDSYPYRYSSTFKIIAVLISNTHPSPLFLSLSVFALHPLYLRVQALSENIP EEIKQEIEKAKQQLDGKEVDYEAAVATKLSIAKKVFNQEKDLILNSSSFQQFFSENEGWLKPYAAFCFLRDFFETSERSQWGRFAQYSEDKLEKLVSTES LHYEIICFHYYVQYHLHLQLSEASEYARKKGVILKGDLPIGVDRNSVDTWVYPNLFRMNTSTGAPPDYFDKNGQNWGFPTYNWEEMSKDNYGWWRARLTQ MGKYFTAYRIDHILGFFRIWELPDHAVTGLVGKFRPSIPLSQEELEKEGIWDFNRLSRPYIRQEILQEKFGSAWAFIATAFLNEYDKNCYEFKEDSNTEK KIVSKLKTSGESSLLLESEDKMRRNLIDLLQNIVLIRDPENPKDFYPRFNLEDTSSFQALDDHSKNVLKRLYYDYYFHRQENLWRQNALKTLPALLNSSE MLACGEDLGLIPSCVHPVCQLYSL | |||
| Full Sequence |
|---|
| Protein Sequence Length: 1023 Download |
| MVNPGLVSGN KPVNSVKISF RLPYLTQWGQ SLLVCGSVPV LGSWNVKKGV LLSPFHEGSE 60 LIWSGSITVP KGFQCEYTYY VVDDKKNIVR WEMGKKHELA LPDGVQSGQE IEFRDLWQTG 120 SDALPFRSAF RDVIFRKSWD SSVKTTTGAN HINLEPEAES ILIQFKVFCP NIEKDTSIYV 180 IGSNTKLGQW KVENGLKLSY VGEFVWLAEC VIQRTLQSSL TIISTYRYCK YGRSGNASIE 240 NGPNREVSIS ASRREAKYIF LSDGMIRETP WRGAGVAIPM FSIRSESDLG VGEFLDLKLL 300 VDWAVASGFH LVQLLPINDT SVHQMWWDSY PYRYSSTFKI IAVLISNTHP SPLFLSLSVF 360 ALHPLYLRVQ ALSENIPEEI KQEIEKAKQQ LDGKEVDYEA AVATKLSIAK KVFNQEKDLI 420 LNSSSFQQFF SENEGWLKPY AAFCFLRDFF ETSERSQWGR FAQYSEDKLE KLVSTESLHY 480 EIICFHYYVQ YHLHLQLSEA SEYARKKGVI LKGDLPIGVD RNSVDTWVYP NLFRMNTSTG 540 APPDYFDKNG QNWGFPTYNW EEMSKDNYGW WRARLTQMGK YFTAYRIDHI LGFFRIWELP 600 DHAVTGLVGK FRPSIPLSQE ELEKEGIWDF NRLSRPYIRQ EILQEKFGSA WAFIATAFLN 660 EYDKNCYEFK EDSNTEKKIV SKLKTSGESS LLLESEDKMR RNLIDLLQNI VLIRDPENPK 720 DFYPRFNLED TSSFQALDDH SKNVLKRLYY DYYFHRQENL WRQNALKTLP ALLNSSEMLA 780 CGEDLGLIPS CVHPVCQLYS LSSNKQEPPS HSCVMQELGL VGLRIQRMPN ESDLEFGIPS 840 QYSYMTVCAP SCHDCSTLRA WWEEDEERRQ RFFKNVMESD ELPPDQCVPE VAHFIIRQHI 900 ESPSMWAIFP LQDLLALKEE YTARPAIEET INDPTNPKHY WRYRVHVTLE SLNKDNELKT 960 IIKDLVRWGG RSVPLEDSQA EANLISTSSV ADTVSEKQQF AGTGEKIRHP SEYNGIPSLT 1020 AR* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
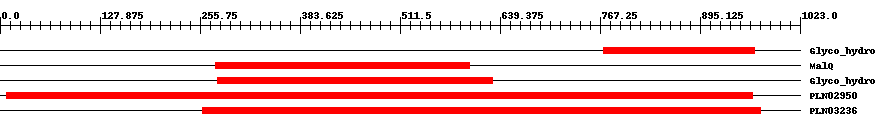 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam02446 | Glyco_hydro_77 | 7.0e-44 | 772 | 965 | 195 | + 4-alpha-glucanotransferase. These enzymes EC:2.4.1.25 transfer a segment of a (1,4)-alpha-D-glucan to a new 4-position in an acceptor, which may be glucose or (1,4)-alpha-D-glucan. | ||
| COG1640 | MalQ | 6.0e-64 | 275 | 601 | 329 | + 4-alpha-glucanotransferase [Carbohydrate transport and metabolism] | ||
| pfam02446 | Glyco_hydro_77 | 2.0e-102 | 278 | 630 | 355 | + 4-alpha-glucanotransferase. These enzymes EC:2.4.1.25 transfer a segment of a (1,4)-alpha-D-glucan to a new 4-position in an acceptor, which may be glucose or (1,4)-alpha-D-glucan. | ||
| PLN02950 | PLN02950 | 0 | 8 | 962 | 957 | + 4-alpha-glucanotransferase | ||
| PLN03236 | PLN03236 | 0 | 259 | 972 | 741 | + 4-alpha-glucanotransferase; Provisional | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0003824 | catalytic activity |
| GO:0004134 | 4-alpha-glucanotransferase activity |
| GO:0005975 | carbohydrate metabolic process |
| GO:2001070 | starch binding |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAR99599.1 | 0 | 1 | 986 | 1 | 934 | 4-alpha-glucanotransferase [Solanum tuberosum] |
| EMBL | CBI32836.1 | 0 | 1 | 1017 | 1 | 960 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002278329.1 | 0 | 1 | 1017 | 1 | 960 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002308854.1 | 0 | 1 | 1017 | 1 | 982 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002323208.1 | 0 | 1 | 983 | 6 | 902 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2x1i_A | 3e-25 | 396 | 648 | 95 | 343 | A Chain A, Glycoside Hydrolase Family 77 4-Alpha-Glucanotransferase From Thermus Brockianus |
| PDB | 1tz7_B | 4e-24 | 272 | 611 | 22 | 329 | A Chain A, Aquifex Aeolicus Amylomaltase |
| PDB | 1tz7_B | 0.003 | 809 | 973 | 346 | 505 | A Chain A, Aquifex Aeolicus Amylomaltase |
| PDB | 1tz7_A | 4e-24 | 272 | 611 | 22 | 329 | A Chain A, Aquifex Aeolicus Amylomaltase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
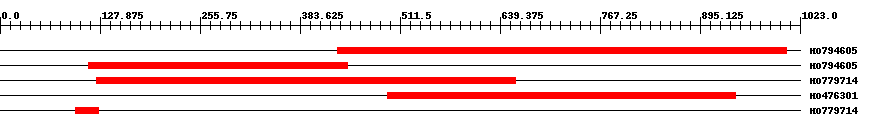 | ||||
| Hit | Length | Start | End | EValue |
| HO794605 | 584 | 432 | 1006 | 0 |
| HO794605 | 333 | 113 | 445 | 0 |
| HO779714 | 536 | 124 | 659 | 0 |
| HO476301 | 449 | 496 | 940 | 0 |
| HO779714 | 31 | 96 | 126 | 0.45 |
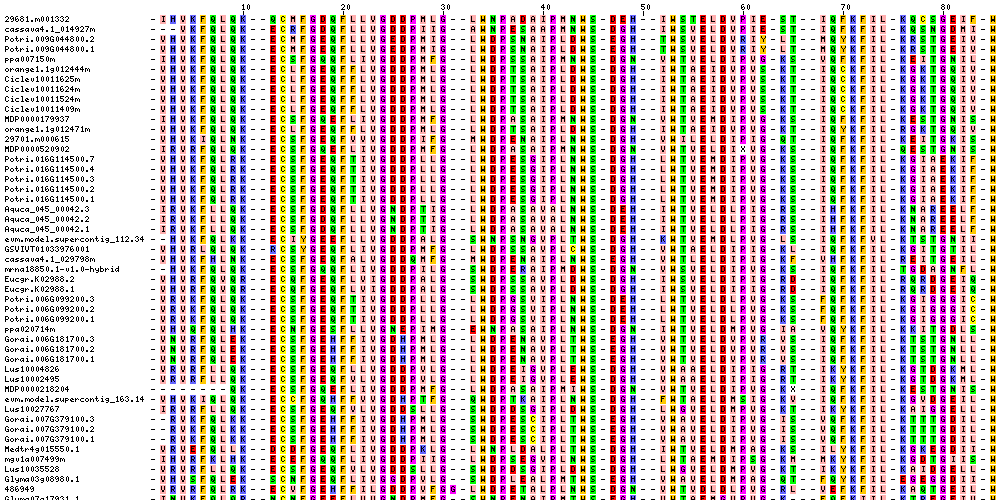
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |