

| Basic Information | |
|---|---|
| Species | Aquilegia coerulea |
| Cazyme ID | Aquca_007_00606.1 |
| Family | CE10 |
| Protein Properties | Length: 602 Molecular Weight: 66282.5 Isoelectric Point: 6.5775 |
| Chromosome | Chromosome/Scaffold: 7 Start: 4857273 End: 4862063 |
| Description | alpha/beta-Hydrolases superfamily protein |
| View CDS | |
| External Links |
|---|
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| CE10 | 32 | 283 | 0 |
| KVPPGFDPKTYVTSKNLLILPDRHVSARLYLPKITNINKKLPVLVYYHGGAFCLSSPSSPMYHNYLNSLVSKANVIAVSVNFRLPPEHRLPIAYEDSWAA LQWVASHSKGQGSESWLAEYADFNKLFLAGDSSGANIAHNMAMLAGDPKTGLSVKILGIALVHPYFWGSKRIGSDTDDPRINPVGDGAPSLLCLGCTRVL VCVAGKDILKDRGWLYYDTLNKCGWNGVVEFLETEGEKHSFHLKKPTGQKAN | |||
| CE10 | 321 | 596 | 0 |
| TGVQSKDVSIQKETGLSARLYIPKIISNDQKLPLLIYFHGGGFFAESAFSPAYHSYLNSVVAEANVVVVSVEYRLAPEHPLPIAYHDSWEAIQWVASHSK GEGHEPWLKDYADFDRLFLAGDSAGANIVHNMALRAGEEELTNGVKILGSILVHPYFWGNEQIGSEGSDPVKNDMINKIWLIACPSSTGLDDPHINPVAK EAPSLSKLGCTRVLVFVAGKDMLRHRGVIYGETLQKSGWGGVVEIIETEGEDHVFHLLNPTCDKAKSFMNNLVSFM | |||
| Full Sequence |
|---|
| Protein Sequence Length: 602 Download |
| MESQKAEVAY EVLPYLRVYQ DGHVERLSET EKVPPGFDPK TYVTSKNLLI LPDRHVSARL 60 YLPKITNINK KLPVLVYYHG GAFCLSSPSS PMYHNYLNSL VSKANVIAVS VNFRLPPEHR 120 LPIAYEDSWA ALQWVASHSK GQGSESWLAE YADFNKLFLA GDSSGANIAH NMAMLAGDPK 180 TGLSVKILGI ALVHPYFWGS KRIGSDTDDP RINPVGDGAP SLLCLGCTRV LVCVAGKDIL 240 KDRGWLYYDT LNKCGWNGVV EFLETEGEKH SFHLKKPTGQ KANVLNERLI STGTSLPVAY 300 EDSLAALKWV AYHSASIDPQ TGVQSKDVSI QKETGLSARL YIPKIISNDQ KLPLLIYFHG 360 GGFFAESAFS PAYHSYLNSV VAEANVVVVS VEYRLAPEHP LPIAYHDSWE AIQWVASHSK 420 GEGHEPWLKD YADFDRLFLA GDSAGANIVH NMALRAGEEE LTNGVKILGS ILVHPYFWGN 480 EQIGSEGSDP VKNDMINKIW LIACPSSTGL DDPHINPVAK EAPSLSKLGC TRVLVFVAGK 540 DMLRHRGVIY GETLQKSGWG GVVEIIETEG EDHVFHLLNP TCDKAKSFMN NLVSFMNHDN 600 K* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
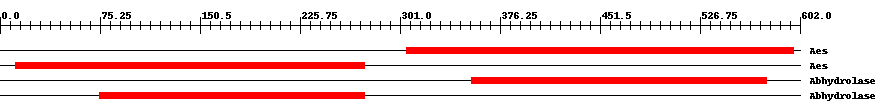 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG0657 | Aes | 1.0e-26 | 306 | 597 | 302 | + Esterase/lipase [Lipid metabolism] | ||
| COG0657 | Aes | 4.0e-27 | 12 | 274 | 292 | + Esterase/lipase [Lipid metabolism] | ||
| pfam07859 | Abhydrolase_3 | 2.0e-41 | 355 | 577 | 227 | + alpha/beta hydrolase fold. This catalytic domain is found in a very wide range of enzymes. | ||
| pfam07859 | Abhydrolase_3 | 9.0e-53 | 75 | 274 | 226 | + alpha/beta hydrolase fold. This catalytic domain is found in a very wide range of enzymes. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0008152 | metabolic process |
| GO:0016787 | hydrolase activity |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI32438.1 | 4.06377e-44 | 23 | 195 | 498 | 663 | unnamed protein product [Vitis vinifera] |
| EMBL | CBI32438.1 | 0 | 60 | 475 | 232 | 663 | unnamed protein product [Vitis vinifera] |
| EMBL | CBI32438.1 | 0 | 340 | 597 | 232 | 437 | unnamed protein product [Vitis vinifera] |
| RefSeq | XP_002284587.1 | 0 | 1 | 291 | 1 | 316 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002462487.1 | 0 | 1 | 596 | 1 | 631 | hypothetical protein SORBIDRAFT_02g026550 [Sorghum bicolor] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2zsi_A | 1.99965e-42 | 314 | 597 | 61 | 349 | A Chain A, Crystal Structure Of The Putative Acetylxylan Esterase From Clostridium Acetobutylicum, Northeast Structural Genomics Target Car6 |
| PDB | 2zsi_A | 1e-29 | 7 | 287 | 35 | 343 | A Chain A, Crystal Structure Of The Putative Acetylxylan Esterase From Clostridium Acetobutylicum, Northeast Structural Genomics Target Car6 |
| PDB | 2zsh_A | 1.99965e-42 | 314 | 597 | 61 | 349 | B Chain B, Structural Basis Of Gibberellin(Ga3)-Induced Della Recognition By The Gibberellin Receptor |
| PDB | 2zsh_A | 1e-29 | 7 | 287 | 35 | 343 | B Chain B, Structural Basis Of Gibberellin(Ga3)-Induced Della Recognition By The Gibberellin Receptor |
| PDB | 2o7v_A | 3e-40 | 314 | 596 | 42 | 327 | B Chain B, Structural Basis Of Gibberellin(Ga3)-Induced Della Recognition By The Gibberellin Receptor |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
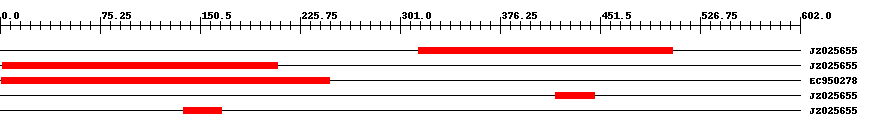 | ||||
| Hit | Length | Start | End | EValue |
| JZ025655 | 192 | 315 | 506 | 0 |
| JZ025655 | 208 | 2 | 209 | 0 |
| EC950278 | 273 | 1 | 248 | 0 |
| JZ025655 | 30 | 418 | 447 | 0.0000000008 |
| JZ025655 | 30 | 138 | 167 | 0.0001 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |