

| Basic Information | |
|---|---|
| Species | Medicago truncatula |
| Cazyme ID | Medtr4g082950.1 |
| Family | GT35 |
| Protein Properties | Length: 886 Molecular Weight: 100750 Isoelectric Point: 4.9334 |
| Chromosome | Chromosome/Scaffold: 4 Start: 28201424 End: 28208121 |
| Description | Glycosyl transferase, family 35 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT35 | 170 | 494 | 0 |
| ALKKLGYNLEDVANQEPDAALGNGGLGRLASCFLDSLATLNYPAWGYGLRYRYGLFKQRITKDGQEEVAENWLEMGNPWEIQRNDVSYPVRFYGEVISGP NEAKQWTGGESILAVAYDVPIPGYKTRTTINLRLWSTKVSTEEFDLKAFNTGDHAKAYAAMKNAEKICYILYPGDESIEGKTLRLKQQYTLCSASLQDII ARFEKRSGMTVNWDSLPDKVVVQMNDTHPTLCIPELIRILIDVKGLSWEKAWDITKRTVAYTNHTVLPEALEKWSLTLLQDLLPRHVEIIKRIDEEFTHE IVSEYGTNDLNMLQEKLGKMRILEN | |||
| GT35 | 578 | 876 | 0 |
| PMMVRMANLCVVGGFSVNGVAEIHSEIVKEEVFNEFYELWPEKFQNKTNGVTPRRWIRFCNPDLSKIITKWIGTEDWVTDLEKLAVLRKFADNEDLQSEW IESKRRNKINVASFIKEKTGYVVSPDAMFDVQVKRIHEYKRQLLNIMGIVYRYKKMKELSAEERKQLFVPRVCIFGGKAFATYVQAKRIVKFITDVGATV NHDPEIGDLLKVIFVPDYNVSVAEMLIPGSELSQHISTAGMEASGTSNMKFAMNGCVQIGTLDGANVEIREEVGEDNFFLFGARAQEIAGLRKERAEGK | |||
| Full Sequence |
|---|
| Protein Sequence Length: 886 Download |
| MSALPFSTTC RHSNSPLHPH SKSNFTGLGQ RNNIWQLFVI TKSNSRRAIR KLYVKNVASD 60 KTAELKERLI KQGTTSNEFV TDSASVASSI KYHAEFTTSF SPEKFEPSKA YFATAESVRD 120 SLIINWNATY EYYERVNVKQ AYYMSMEYLQ GRALLNAIGN LQLSGPYAEA LKKLGYNLED 180 VANQEPDAAL GNGGLGRLAS CFLDSLATLN YPAWGYGLRY RYGLFKQRIT KDGQEEVAEN 240 WLEMGNPWEI QRNDVSYPVR FYGEVISGPN EAKQWTGGES ILAVAYDVPI PGYKTRTTIN 300 LRLWSTKVST EEFDLKAFNT GDHAKAYAAM KNAEKICYIL YPGDESIEGK TLRLKQQYTL 360 CSASLQDIIA RFEKRSGMTV NWDSLPDKVV VQMNDTHPTL CIPELIRILI DVKGLSWEKA 420 WDITKRTVAY TNHTVLPEAL EKWSLTLLQD LLPRHVEIIK RIDEEFTHEI VSEYGTNDLN 480 MLQEKLGKMR ILENIELPDS VVEFINNTVL ADDPVEEIDV DDNDIKATEK KDNEEENDDD 540 DEEEEDEVGK DEQEADDGEE LVVENKKEWK FEVDPNLPMM VRMANLCVVG GFSVNGVAEI 600 HSEIVKEEVF NEFYELWPEK FQNKTNGVTP RRWIRFCNPD LSKIITKWIG TEDWVTDLEK 660 LAVLRKFADN EDLQSEWIES KRRNKINVAS FIKEKTGYVV SPDAMFDVQV KRIHEYKRQL 720 LNIMGIVYRY KKMKELSAEE RKQLFVPRVC IFGGKAFATY VQAKRIVKFI TDVGATVNHD 780 PEIGDLLKVI FVPDYNVSVA EMLIPGSELS QHISTAGMEA SGTSNMKFAM NGCVQIGTLD 840 GANVEIREEV GEDNFFLFGA RAQEIAGLRK ERAEGKASFK SKHNI* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
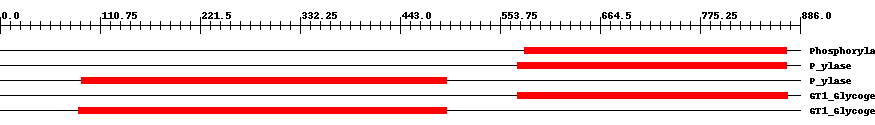 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00343 | Phosphorylase | 2.0e-155 | 581 | 871 | 291 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| TIGR02093 | P_ylase | 4.0e-158 | 573 | 871 | 299 | + glycogen/starch/alpha-glucan phosphorylases. This family consists of phosphorylases. Members use phosphate to break alpha 1,4 linkages between pairs of glucose residues at the end of long glucose polymers, releasing alpha-D-glucose 1-phosphate. The nomenclature convention is to preface the name according to the natural substrate, as in glycogen phosphorylase, starch phosphorylase, maltodextrin phosphorylase, etc. Name differences among these substrates reflect differences in patterns of branching with alpha 1,6 linkages. Members include allosterically regulated and unregulated forms. A related family, TIGR02094, contains examples known to act well on particularly small alpha 1,4 glucans, as may be found after import from exogenous sources [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| TIGR02093 | P_ylase | 9.0e-170 | 90 | 494 | 408 | + glycogen/starch/alpha-glucan phosphorylases. This family consists of phosphorylases. Members use phosphate to break alpha 1,4 linkages between pairs of glucose residues at the end of long glucose polymers, releasing alpha-D-glucose 1-phosphate. The nomenclature convention is to preface the name according to the natural substrate, as in glycogen phosphorylase, starch phosphorylase, maltodextrin phosphorylase, etc. Name differences among these substrates reflect differences in patterns of branching with alpha 1,6 linkages. Members include allosterically regulated and unregulated forms. A related family, TIGR02094, contains examples known to act well on particularly small alpha 1,4 glucans, as may be found after import from exogenous sources [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 1.0e-177 | 573 | 872 | 300 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 87 | 494 | 412 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004645 | phosphorylase activity |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI27267.1 | 0 | 1 | 876 | 1 | 828 | unnamed protein product [Vitis vinifera] |
| Swiss-Prot | P53535 | 0 | 58 | 876 | 54 | 869 | PHSL2_SOLTU RecName: Full=Alpha-1,4 glucan phosphorylase L-2 isozyme, chloroplastic/amyloplastic; AltName: Full=Starch phosphorylase L-2; Flags: Precursor |
| RefSeq | XP_002274575.1 | 0 | 1 | 876 | 1 | 876 | PREDICTED: hypothetical protein [Vitis vinifera] |
| RefSeq | XP_002316098.1 | 0 | 1 | 876 | 1 | 848 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002512108.1 | 0 | 1 | 876 | 1 | 868 | glycogen phosphorylase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 1ygp_B | 0 | 93 | 482 | 51 | 453 | A Chain A, Phosphorylated Form Of Yeast Glycogen Phosphorylase With Phosphate Bound In The Active Site. |
| PDB | 1ygp_B | 0 | 581 | 869 | 468 | 770 | A Chain A, Phosphorylated Form Of Yeast Glycogen Phosphorylase With Phosphate Bound In The Active Site. |
| PDB | 1ygp_A | 0 | 93 | 482 | 51 | 453 | A Chain A, Phosphorylated Form Of Yeast Glycogen Phosphorylase With Phosphate Bound In The Active Site. |
| PDB | 1ygp_A | 0 | 581 | 869 | 468 | 770 | A Chain A, Phosphorylated Form Of Yeast Glycogen Phosphorylase With Phosphate Bound In The Active Site. |
| PDB | 1z8d_A | 0 | 64 | 493 | 9 | 433 | A Chain A, Crystal Structure Of Human Muscle Glycogen Phosphorylase A With Amp And Glucose |
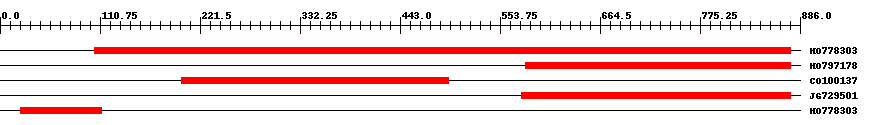
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| HO778303 | 773 | 105 | 876 | 0 |
| HO797178 | 295 | 582 | 876 | 0 |
| CO100137 | 297 | 201 | 497 | 0 |
| JG729501 | 298 | 578 | 875 | 0 |
| HO778303 | 90 | 23 | 112 | 0.01 |
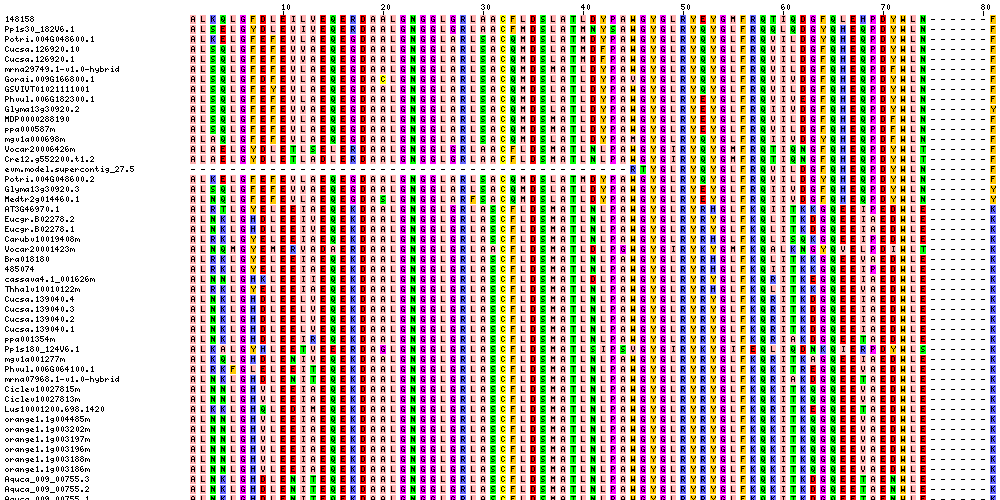
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |