

| Basic Information | |
|---|---|
| Species | Physcomitrella patens |
| Cazyme ID | Pp1s53_212V6.1 |
| Family | GT35 |
| Protein Properties | Length: 988 Molecular Weight: 110565 Isoelectric Point: 7.1029 |
| Chromosome | Chromosome/Scaffold: 53 Start: 1437700 End: 1443949 |
| Description | Glycosyl transferase, family 35 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| Plaza |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT35 | 179 | 499 | 0 |
| ALRKLGHDLEAVAEQEPDAALGNGGLGRLASCFLDSLATLNYPAWGYGLRYKYGLFQQTITKEGQQEQCEKWLEIGYPWEIPRNDISYSIKFFGEVVDSE DGKKKWVGGENVSALAYDVPIPGFRTKNTISLRLWSTRVSAEDFNLEAFNSGDYGKADEAQANAERICYVLYPGDATEEGKLLRLKQQYTLCSASIQDII ARFKERSGGEVNWNAFPTKVAIQMNDTHPTLCVPELLRILIDEEGLSWDQAWKITQATVAYTNHTVLPEALEKWPLIAFQKLLPRHVQIIETIDEQFMKF VASKVDKTELEAKIASMRILE | |||
| GT35 | 566 | 981 | 0 |
| IEEEKEEEVDVEVAPQKLAGTVRMANLCVIAGHKVNGVAAIHSEIVKEEVFNDFYKIWPEKFQNKTNGVTPRRWLNFCNPELSAVITKWLGSEEWVLDTK QLAGLRNFADNQELQKDWVQAKIARKEKLAAYIKTKTGYTINPNALFDIQVKRIHEYKRQLLNIMGVIYRYQNMKKMTPKERAAKYVPRVVMFGGKAFAT YWQAKRIVKLITDVAATINNDPDIGDLLKVIMVPDYNVSVAEVLIPASELSQHISTAGMEASGTSNMKFSMNGCVLIGTLDGANVEIREEVGEDNFFLFG ARATEIAGLRADREAGKFVPDPRFEEVKKFIRSGAFGDYDYSELLGALEGNSGYGRGDYFLVGYDFPSYIECQDKVDEAYRDQERWTRMSIMNTAGSYTF SSDRTIHEYAKDIWDI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 988 Download |
| MASLTVVPSS SSASCSRPTT SPRAATTPKP ALSSASRLRT NTRSLRLLAV PGALKRTPQR 60 KRTCEARAIE TSVSENAETE ILPKANPLST DPNAIASNIK YHAEFTPSFS PYKFELKQAY 120 VATAESLRDT LIQRWNETYK HFTRENAKTI HYLSMEFLQG RALLNAVGNL ELKDAYSEAL 180 RKLGHDLEAV AEQEPDAALG NGGLGRLASC FLDSLATLNY PAWGYGLRYK YGLFQQTITK 240 EGQQEQCEKW LEIGYPWEIP RNDISYSIKF FGEVVDSEDG KKKWVGGENV SALAYDVPIP 300 GFRTKNTISL RLWSTRVSAE DFNLEAFNSG DYGKADEAQA NAERICYVLY PGDATEEGKL 360 LRLKQQYTLC SASIQDIIAR FKERSGGEVN WNAFPTKVAI QMNDTHPTLC VPELLRILID 420 EEGLSWDQAW KITQATVAYT NHTVLPEALE KWPLIAFQKL LPRHVQIIET IDEQFMKFVA 480 SKVDKTELEA KIASMRILEN IDLPGSIQSL LPPKPKKEKV KKTSEPATKV DGGKSFLSAV 540 KELVGTAKSA VKELVGTAKS AVKKVIEEEK EEEVDVEVAP QKLAGTVRMA NLCVIAGHKV 600 NGVAAIHSEI VKEEVFNDFY KIWPEKFQNK TNGVTPRRWL NFCNPELSAV ITKWLGSEEW 660 VLDTKQLAGL RNFADNQELQ KDWVQAKIAR KEKLAAYIKT KTGYTINPNA LFDIQVKRIH 720 EYKRQLLNIM GVIYRYQNMK KMTPKERAAK YVPRVVMFGG KAFATYWQAK RIVKLITDVA 780 ATINNDPDIG DLLKVIMVPD YNVSVAEVLI PASELSQHIS TAGMEASGTS NMKFSMNGCV 840 LIGTLDGANV EIREEVGEDN FFLFGARATE IAGLRADREA GKFVPDPRFE EVKKFIRSGA 900 FGDYDYSELL GALEGNSGYG RGDYFLVGYD FPSYIECQDK VDEAYRDQER WTRMSIMNTA 960 GSYTFSSDRT IHEYAKDIWD IMPSPVA* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
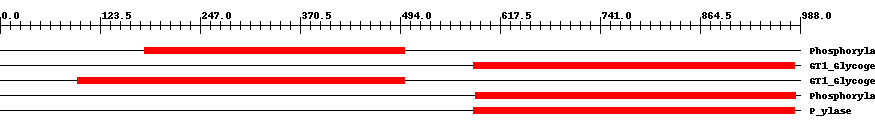 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00343 | Phosphorylase | 3.0e-138 | 179 | 499 | 321 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 585 | 981 | 402 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 96 | 500 | 409 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| pfam00343 | Phosphorylase | 0 | 587 | 983 | 402 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| TIGR02093 | P_ylase | 0 | 585 | 981 | 402 | + glycogen/starch/alpha-glucan phosphorylases. This family consists of phosphorylases. Members use phosphate to break alpha 1,4 linkages between pairs of glucose residues at the end of long glucose polymers, releasing alpha-D-glucose 1-phosphate. The nomenclature convention is to preface the name according to the natural substrate, as in glycogen phosphorylase, starch phosphorylase, maltodextrin phosphorylase, etc. Name differences among these substrates reflect differences in patterns of branching with alpha 1,6 linkages. Members include allosterically regulated and unregulated forms. A related family, TIGR02094, contains examples known to act well on particularly small alpha 1,4 glucans, as may be found after import from exogenous sources [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004645 | phosphorylase activity |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| Swiss-Prot | P04045 | 0 | 54 | 987 | 33 | 966 | PHSL1_SOLTU RecName: Full=Alpha-1,4 glucan phosphorylase L-1 isozyme, chloroplastic/amyloplastic; AltName: Full=Starch phosphorylase L-1; Flags: Precursor |
| Swiss-Prot | P27598 | 0 | 39 | 983 | 15 | 951 | PHSL_IPOBA RecName: Full=Alpha-1,4 glucan phosphorylase L isozyme, chloroplastic/amyloplastic; AltName: Full=Starch phosphorylase L; Flags: Precursor |
| RefSeq | XP_001762173.1 | 0 | 1 | 987 | 1 | 923 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_001775705.1 | 0 | 60 | 986 | 71 | 974 | predicted protein [Physcomitrella patens subsp. patens] |
| RefSeq | XP_002274575.1 | 0 | 86 | 983 | 98 | 977 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2azd_B | 0 | 540 | 979 | 360 | 792 | B Chain B, Reassembly And Co-Crystallization Of A Family 9 Processive Endoglucanase From Separately Expressed Gh9 And Cbm3c Modules |
| PDB | 2azd_B | 0 | 103 | 479 | 27 | 382 | B Chain B, Reassembly And Co-Crystallization Of A Family 9 Processive Endoglucanase From Separately Expressed Gh9 And Cbm3c Modules |
| PDB | 2azd_A | 0 | 540 | 979 | 360 | 792 | B Chain B, Reassembly And Co-Crystallization Of A Family 9 Processive Endoglucanase From Separately Expressed Gh9 And Cbm3c Modules |
| PDB | 2azd_A | 0 | 103 | 479 | 27 | 382 | B Chain B, Reassembly And Co-Crystallization Of A Family 9 Processive Endoglucanase From Separately Expressed Gh9 And Cbm3c Modules |
| PDB | 2aw3_B | 0 | 540 | 979 | 360 | 792 | B Chain B, Reassembly And Co-Crystallization Of A Family 9 Processive Endoglucanase From Separately Expressed Gh9 And Cbm3c Modules |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| starch degradation I | RXN-1826 | EC-2.4.1.1 | phosphorylase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
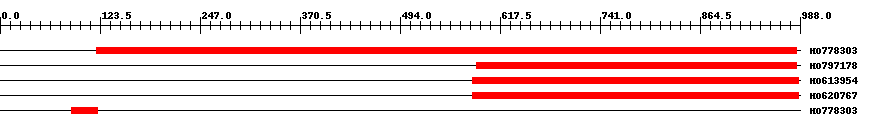 | ||||
| Hit | Length | Start | End | EValue |
| HO778303 | 869 | 119 | 984 | 0 |
| HO797178 | 397 | 588 | 984 | 0 |
| HO613954 | 403 | 584 | 986 | 0 |
| HO620767 | 403 | 584 | 986 | 0 |
| HO778303 | 34 | 88 | 121 | 0.001 |
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
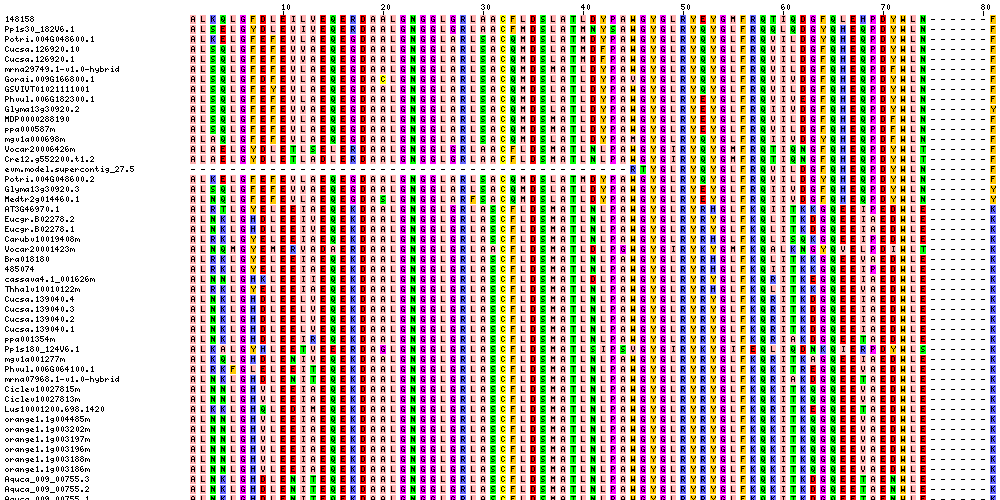 |