

| Basic Information | |
|---|---|
| Species | Prunus persica |
| Cazyme ID | ppa000827m |
| Family | GT35 |
| Protein Properties | Length: 990 Molecular Weight: 112163 Isoelectric Point: 4.8906 |
| Chromosome | Chromosome/Scaffold: 3 Start: 19171126 End: 19177833 |
| Description | Glycosyl transferase, family 35 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT35 | 171 | 506 | 0 |
| ALSKLGHKLENVACQEPDAALGNGGLGRLASCFLDSLATLNYPAWGYGLRYKYGLFKQRITKDGQEEVAEDWLELGNPWEIVRNDVSYPIKFYGKVVSGS DGKRHWIGGEDIDAVAYDVPIPGYKTKTTINLRLWSTKASSQDFDLYAFNSGEHTKASEALANAEKICYVLYPGDESVEGKTLRLKQQYTLCSASLQDIV ERFERRSGPNIKWEEFPEKVAVQMNDTHPTLCIPELMRILIDLKGLSWKEAWNITQRTVAYTNHTVLPEALEKWSLELMQKLLPRHVEIIEMIDEELINT IILEYGTADYDLLEKKLKEMRILENVDLPATFADLF | |||
| GT35 | 586 | 983 | 0 |
| PKLVRMANLCVVGGHAVNGVAEIHSEIVKDEVFNSFFKLWPKKFQNKTNGVTPRRWIRFCNPDLSKIITKWIGTEDWVLNTENLAELRKFADNNDLQTQW REAKRSNKLKVVSLIKERTGYSVSPDAMFDIQVKRIHEYKRQLLNIFGIVYRYKKMKEMSASGRKAKFVPRVCMFGGKAFSTYVQAKRIVKFITDVAATI NRDPGIGDLLKVVFVPDYNVSVAELLIPASELSQHISTAGMEASGTSNMKFAMNGCVLIGTLDGANVEIREEVGADNFFLFGAKAHEIAGLRKERAEGKF VPDPRFEEVKEFIRSGVFGSFNYDELIGSLEGNEGFGRADYFLVGKDFPSYIECQEKVDEAYRDQKRWTRMSILNTAGSYKFSSDRTIHEYAEDIWNI | |||
| Full Sequence |
|---|
| Protein Sequence Length: 990 Download |
| MAASQFAATR GGAETVWQCK SQSKLIDFSS RKNKSKLLFT RRNLNQRRSF SFSVKNASNE 60 SSQKLKDPIV EQDSSILSSF IPDAASIASS IKYHAEFTAS FSPERFELPK AFFATAQSVR 120 DALIINWNAT YAYYEKLNAK QAYYLSMEFL QGRALLNAIG NLELDGAYAE ALSKLGHKLE 180 NVACQEPDAA LGNGGLGRLA SCFLDSLATL NYPAWGYGLR YKYGLFKQRI TKDGQEEVAE 240 DWLELGNPWE IVRNDVSYPI KFYGKVVSGS DGKRHWIGGE DIDAVAYDVP IPGYKTKTTI 300 NLRLWSTKAS SQDFDLYAFN SGEHTKASEA LANAEKICYV LYPGDESVEG KTLRLKQQYT 360 LCSASLQDIV ERFERRSGPN IKWEEFPEKV AVQMNDTHPT LCIPELMRIL IDLKGLSWKE 420 AWNITQRTVA YTNHTVLPEA LEKWSLELMQ KLLPRHVEII EMIDEELINT IILEYGTADY 480 DLLEKKLKEM RILENVDLPA TFADLFVKPK ESSVVVPSEE LEDSKEEEEE DESVDEENES 540 VDEEDESVDE EDESVDEEDE SVDEENGPDK KCDEEKKKKV VVEPPPKLVR MANLCVVGGH 600 AVNGVAEIHS EIVKDEVFNS FFKLWPKKFQ NKTNGVTPRR WIRFCNPDLS KIITKWIGTE 660 DWVLNTENLA ELRKFADNND LQTQWREAKR SNKLKVVSLI KERTGYSVSP DAMFDIQVKR 720 IHEYKRQLLN IFGIVYRYKK MKEMSASGRK AKFVPRVCMF GGKAFSTYVQ AKRIVKFITD 780 VAATINRDPG IGDLLKVVFV PDYNVSVAEL LIPASELSQH ISTAGMEASG TSNMKFAMNG 840 CVLIGTLDGA NVEIREEVGA DNFFLFGAKA HEIAGLRKER AEGKFVPDPR FEEVKEFIRS 900 GVFGSFNYDE LIGSLEGNEG FGRADYFLVG KDFPSYIECQ EKVDEAYRDQ KRWTRMSILN 960 TAGSYKFSSD RTIHEYAEDI WNINPVELP* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
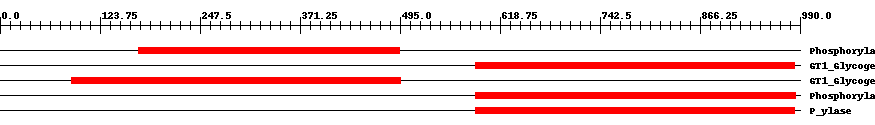
| pfam00343 | Phosphorylase | 1.0e-135 | 171 | 494 | 324 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 589 | 983 | 400 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| cd04300 | GT1_Glycogen_Phosphorylase | 0 | 88 | 495 | 412 | + This is a family of oligosaccharide phosphorylases. It includes yeast and mammalian glycogen phosphorylases, plant starch/glucan phosphorylase, as well as the maltodextrin phosphorylases of bacteria. The members of this family catalyze the breakdown of oligosaccharides into glucose-1-phosphate units. They are important allosteric enzymes in carbohydrate metabolism. The allosteric control mechanisms of yeast and mammalian members of this family are different from that of bacterial members. The members of this family belong to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. | ||
| pfam00343 | Phosphorylase | 0 | 589 | 985 | 402 | + Carbohydrate phosphorylase. The members of this family catalyze the formation of glucose 1-phosphate from one of the following polyglucoses; glycogen, starch, glucan or maltodextrin. | ||
| TIGR02093 | P_ylase | 0 | 589 | 983 | 400 | + glycogen/starch/alpha-glucan phosphorylases. This family consists of phosphorylases. Members use phosphate to break alpha 1,4 linkages between pairs of glucose residues at the end of long glucose polymers, releasing alpha-D-glucose 1-phosphate. The nomenclature convention is to preface the name according to the natural substrate, as in glycogen phosphorylase, starch phosphorylase, maltodextrin phosphorylase, etc. Name differences among these substrates reflect differences in patterns of branching with alpha 1,6 linkages. Members include allosterically regulated and unregulated forms. A related family, TIGR02094, contains examples known to act well on particularly small alpha 1,4 glucans, as may be found after import from exogenous sources [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004645 | phosphorylase activity |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CBI22291.1 | 0 | 21 | 989 | 23 | 982 | unnamed protein product [Vitis vinifera] |
| Swiss-Prot | P04045 | 0 | 8 | 988 | 2 | 965 | PHSL1_SOLTU RecName: Full=Alpha-1,4 glucan phosphorylase L-1 isozyme, chloroplastic/amyloplastic; AltName: Full=Starch phosphorylase L-1; Flags: Precursor |
| Swiss-Prot | P53536 | 0 | 31 | 988 | 39 | 1002 | PHSL_VICFA RecName: Full=Alpha-1,4 glucan phosphorylase L isozyme, chloroplastic/amyloplastic; AltName: Full=Starch phosphorylase L; Flags: Precursor |
| RefSeq | XP_002305367.1 | 0 | 42 | 989 | 4 | 949 | predicted protein [Populus trichocarpa] |
| RefSeq | XP_002526085.1 | 0 | 17 | 989 | 15 | 977 | glycogen phosphorylase, putative [Ricinus communis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2azd_B | 0 | 587 | 981 | 402 | 792 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2azd_B | 0 | 138 | 464 | 57 | 375 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2azd_A | 0 | 587 | 981 | 402 | 792 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2azd_A | 0 | 138 | 464 | 57 | 375 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| PDB | 2aw3_B | 0 | 587 | 981 | 402 | 792 | A Chain A, Crystal Structure Of A Family Gt4 Glycosyltransferase From Bacillus Anthracis Orf Ba1558. |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
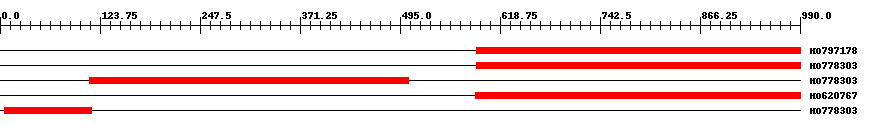 | ||||
| Hit | Length | Start | End | EValue |
| HO797178 | 401 | 590 | 990 | 0 |
| HO778303 | 401 | 590 | 990 | 0 |
| HO778303 | 396 | 111 | 506 | 0 |
| HO620767 | 403 | 588 | 990 | 0 |
| HO778303 | 118 | 5 | 113 | 0.00000000000004 |
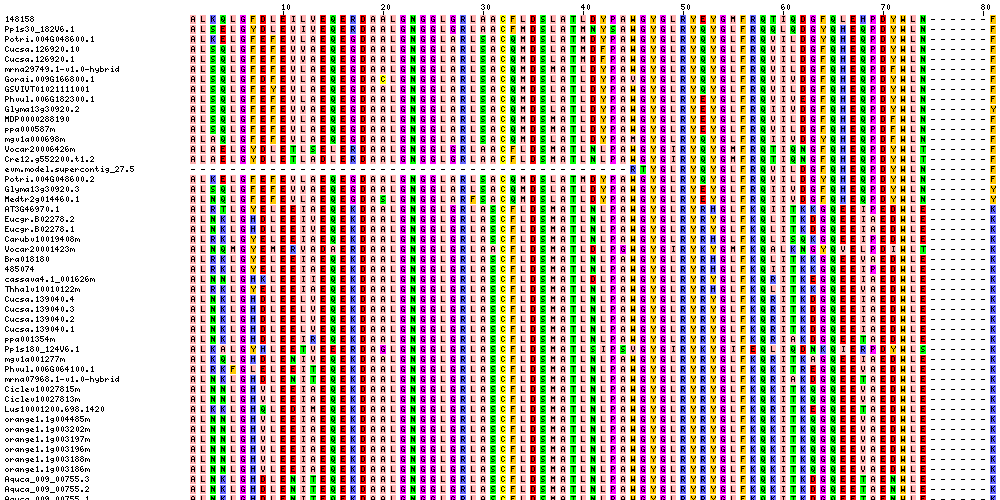
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |