

| Basic Information | |
|---|---|
| Species | Brassica rapa |
| Cazyme ID | Bra022797 |
| Family | GH19 |
| Protein Properties | Length: 145 Molecular Weight: 15956.9 Isoelectric Point: 7.2656 |
| Chromosome | Chromosome/Scaffold: 03 Start: 7107086 End: 7107884 |
| Description | Chitinase family protein |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH19 | 1 | 144 | 0 |
| MFAHFTHETGHFCYIEEINGASRDYCDENNRQYPCAPGKGYYGRGPIQLSWNYNYGPCGQSLGLNLLGQPELVGSNPTVTFRTALWFWMNSVRPVLNQGF GATIRAINGMECNSGNPAAVNARIMYYRNYCGQLGVDPGPNLGC | |||
| Full Sequence |
|---|
| Protein Sequence Length: 145 Download |
| MFAHFTHETG HFCYIEEING ASRDYCDENN RQYPCAPGKG YYGRGPIQLS WNYNYGPCGQ 60 SLGLNLLGQP ELVGSNPTVT FRTALWFWMN SVRPVLNQGF GATIRAINGM ECNSGNPAAV 120 NARIMYYRNY CGQLGVDPGP NLGC* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG3158 | Kup | 0.0004 | 40 | 77 | 38 | + K+ transporter [Inorganic ion transport and metabolism] | ||
| COG3179 | COG3179 | 8.0e-5 | 34 | 88 | 57 | + Predicted chitinase [General function prediction only] | ||
| cd00442 | lysozyme_like | 2.0e-12 | 34 | 94 | 61 | + lysozyme_like domain. This contains several members including Soluble Lytic Transglycosylases (SLT), Goose Egg-White Lysozymes (GEWL), Hen Egg-White Lysozymes (HEWL), chitinases, bacteriophage lambda lysozymes, endolysins, autolysins, and chitosanases. All the members are involved in the hydrolysis of beta-1,4- linked polysaccharides. | ||
| pfam00182 | Glyco_hydro_19 | 2.0e-74 | 1 | 144 | 177 | + Chitinase class I. | ||
| cd00325 | chitinase_glyco_hydro_19 | 1.0e-74 | 1 | 144 | 176 | + Glycoside hydrolase family 19 chitinase domain. Chitinases are enzymes that catalyze the hydrolysis of the beta-1,4-N-acetyl-D-glucosamine linkages in chitin polymers. Family 19 chitinases are found primarily in plants (classes I, III, and IV), but some are found in bacteria. Class I and II chitinases are similar in their catalytic domains. Class I chitinases have an N-terminal cysteine-rich, chitin-binding domain which is separated from the catalytic domain by a proline and glycine-rich hinge region. Class II chitinases lack both the chitin-binding domain and the hinge region. Class IV chitinases are similar to class I chitinases but they are smaller in size due to certain deletions. Despite any significant sequence homology with lysozymes, structural analysis reveals that family 19 chitinases, together with family 46 chitosanases, are similar to several lysozymes including those from T4-phage and from goose. The structures reveal that the different enzyme groups arose from a common ancestor glycohydrolase antecedent to the procaryotic/eucaryotic divergence. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004568 | chitinase activity |
| GO:0006032 | chitin catabolic process |
| GO:0016998 | cell wall macromolecule catabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABV89613.1 | 0 | 1 | 144 | 125 | 268 | chitinase [Brassica rapa] |
| DDBJ | BAF35569.1 | 0 | 1 | 144 | 125 | 268 | chitinase [Brassica rapa subsp. pekinensis] |
| RefSeq | NP_181886.1 | 0 | 1 | 144 | 122 | 265 | chitinase, putative [Arabidopsis thaliana] |
| RefSeq | NP_181887.1 | 0 | 1 | 144 | 121 | 264 | chitinase, putative [Arabidopsis thaliana] |
| Swiss-Prot | Q06209 | 0 | 1 | 144 | 125 | 268 | CHI4_BRANA RecName: Full=Basic endochitinase CHB4; Flags: Precursor |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3hbh_A | 0 | 1 | 144 | 59 | 204 | A Chain A, Crystal Structure Of A Rice Os3bglu6 Beta-glucosidase |
| PDB | 3hbe_X | 0 | 1 | 144 | 59 | 204 | A Chain A, Crystal Structure Of A Rice Os3bglu6 Beta-glucosidase |
| PDB | 3hbd_A | 0 | 1 | 144 | 59 | 204 | A Chain A, Class Iv Chitinase Structure From Picea Abies At 1.8a |
| PDB | 2z38_A | 9.00054e-42 | 12 | 144 | 87 | 240 | A Chain A, Crystal Structure Of Chloride Bound Brassica Juncea Chitinase Catalytic Module (Bjchi3) |
| PDB | 2z37_D | 9.00054e-42 | 12 | 144 | 84 | 237 | A Chain A, Crystal Structure Of Brassica Juncea Chitinase Catalytic Module (Bjchi3) |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| chitin degradation II | 3.2.1.14-RXN | EC-3.2.1.14 | chitinase |
| chitin degradation II | RXN-12623 | EC-3.2.1.14 | chitinase |
| chitin degradation II | RXN-12624 | EC-3.2.1.14 | chitinase |
| chitin degradation III (carnivorous plants) | 3.2.1.14-RXN | EC-3.2.1.14 | chitinase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| FD975419 | 145 | 1 | 145 | 0 |
| EY926575 | 145 | 1 | 145 | 0 |
| EY943317 | 145 | 1 | 145 | 0 |
| EE418115 | 145 | 1 | 145 | 0 |
| FY069380 | 145 | 1 | 145 | 0 |
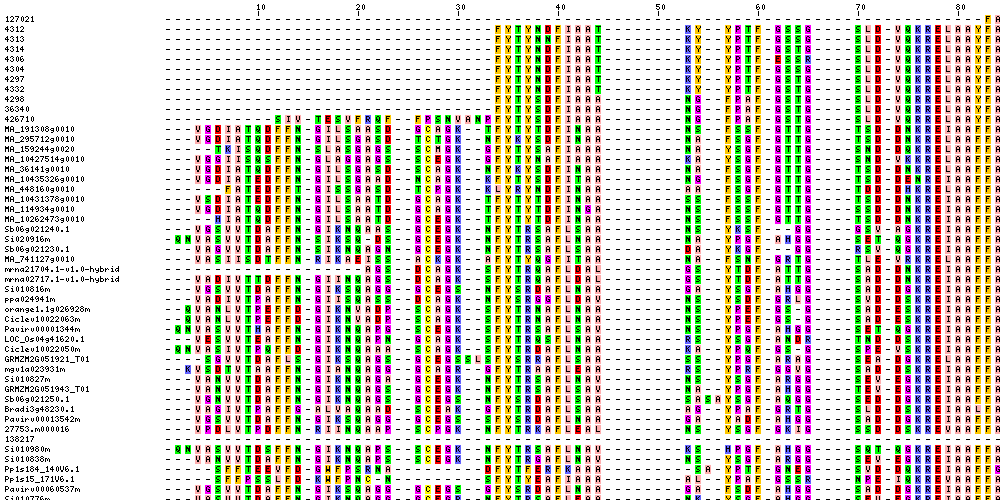
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |