

| Basic Information | |
|---|---|
| Species | Picea abies |
| Cazyme ID | MA_123187g0010 |
| Family | GH19 |
| Protein Properties | Length: 170 Molecular Weight: 18619.8 Isoelectric Point: 7.5069 |
| View CDS | |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
| Family | Start | End | Evalue |
| GH19 | 77 | 159 | 1.1e-23 |
| GSLQFRGNANYGAARTYLGSDLLNSAGLMAQDHLTSWKITLWFWNSNCHTAITSGQGFGPTIQAINGAIECNGENTDPVNDSM | |||
| Full Sequence |
|---|
| Protein Sequence Length: 170 Download |
| MGGNRTWPCE YIVSVDISLG HASGTLNRID LSWGALNVAD RPLRQEGSNL SVLRGGDSQE 60 QLVRFEQHTI SMRIRTGSLQ FRGNANYGAA RTYLGSDLLN SAGLMAQDHL TSWKITLWFW 120 NSNCHTAITS GQGFGPTIQA INGAIECNGE NTDPVNDSMH SVSCRPRKQP |
| Functional Domains Download unfiltered results here | ||||||
|---|---|---|---|---|---|---|
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description |
| pfam00182 | Glyco_hydro_19 | 8.0e-16 | 77 | 158 | 101 | + Chitinase class I. |
| cd00325 | chitinase_glyco_hydro_19 | 3.0e-23 | 77 | 157 | 100 | + Glycoside hydrolase family 19 chitinase domain. Chitinases are enzymes that catalyze the hydrolysis of the beta-1,4-N-acetyl-D-glucosamine linkages in chitin polymers. Family 19 chitinases are found primarily in plants (classes I, III, and IV), but some are found in bacteria. Class I and II chitinases are similar in their catalytic domains. Class I chitinases have an N-terminal cysteine-rich, chitin-binding domain which is separated from the catalytic domain by a proline and glycine-rich hinge region. Class II chitinases lack both the chitin-binding domain and the hinge region. Class IV chitinases are similar to class I chitinases but they are smaller in size due to certain deletions. Despite any significant sequence homology with lysozymes, structural analysis reveals that family 19 chitinases, together with family 46 chitosanases, are similar to several lysozymes including those from T4-phage and from goose. The structures reveal that the different enzyme groups arose from a common ancestor glycohydrolase antecedent to the procaryotic/eucaryotic divergence. |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | ABK21702.1 | 4e-36 | 77 | 159 | 127 | 211 | unknown [Picea sitchensis] |
| GenBank | ABK21750.1 | 2e-35 | 77 | 159 | 126 | 210 | unknown [Picea sitchensis] |
| GenBank | ABK23371.1 | 8e-36 | 77 | 159 | 126 | 210 | unknown [Picea sitchensis] |
| GenBank | ABR17294.1 | 4e-35 | 77 | 159 | 127 | 211 | unknown [Picea sitchensis] |
| GenBank | ACN39919.1 | 2e-35 | 77 | 159 | 127 | 211 | unknown [Picea sitchensis] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3hbh_A | 1e-18 | 77 | 160 | 101 | 185 | A Chain A, High Resolution Structure Of E.coli Wrba With Fmn |
| PDB | 3hbe_X | 1e-18 | 77 | 160 | 101 | 185 | A Chain A, High Resolution Structure Of E.coli Wrba With Fmn |
| PDB | 3hbd_A | 1e-18 | 77 | 160 | 101 | 185 | A Chain A, Class Iv Chitinase Structure From Picea Abies At 1.8a |
| PDB | 2cjl_B | 0.00000000000002 | 77 | 151 | 95 | 176 | A Chain A, Crystal Structure And Enzymatic Properties Of A Bacterial Family 19 Chitinase Reveal Differences With Plant Enzymes |
| PDB | 2cjl_A | 0.00000000000002 | 77 | 151 | 95 | 176 | A Chain A, Crystal Structure And Enzymatic Properties Of A Bacterial Family 19 Chitinase Reveal Differences With Plant Enzymes |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
| Hit | Length | Start | End | EValue |
| DR473845 | 85 | 77 | 159 | 1e-39 |
| DR473845 | 40 | 43 | 80 | 1e-39 |
| DR473845 | 12 | 159 | 170 | 1e-39 |
| DR475002 | 85 | 77 | 159 | 1e-39 |
| DR475002 | 40 | 43 | 80 | 1e-39 |
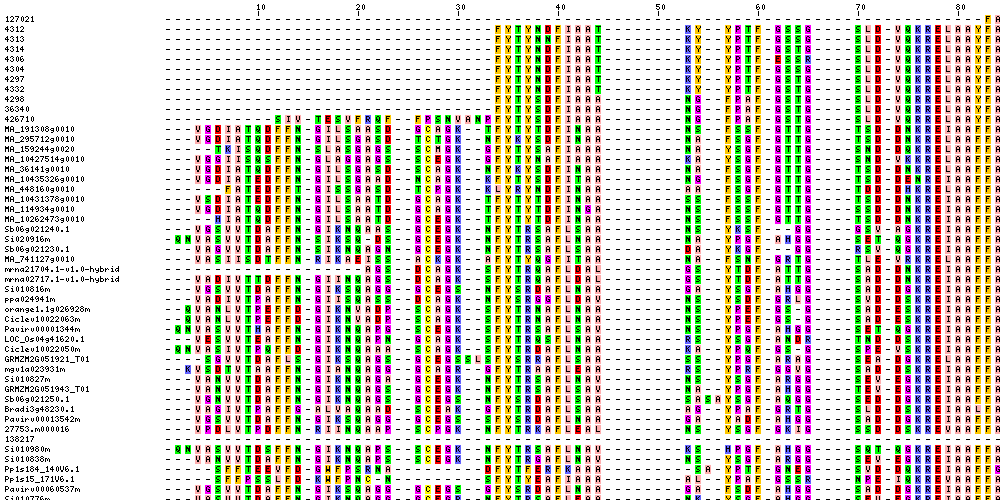
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |