

| Basic Information | |
|---|---|
| Species | Malus domestica |
| Cazyme ID | MDP0000298027 |
| Family | GH17 |
| Protein Properties | Length: 480 Molecular Weight: 53342.2 Isoelectric Point: 4.3405 |
| Chromosome | Chromosome/Scaffold: 015030647 Start: 1086 End: 4669 |
| Description | beta-1,3-glucanase 4 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 167 | 479 | 0 |
| IGVCYGMIGNDLPSATEVVNLYKTYGIGKMRLFDRNPSALQALRGRQINVTLGIRNEDLPNLAVSQDAVNSWFASNVEPYLNDIIFNYISVGNEVIPGAL GDYVLPVMLSLQNILDEKNLACIKVTTVVPGSALGVSYPPSSGEFTSEVSSIMSGIVPFLAQQSSPLMINVYPYFAYASDPANIRLDYAQFTATSPVVQD GSLSYYNMFDAMVDAFLAAMEKIDGVGANVVDVVVSESGWPSDGNGNFTTPGLAATYNKNYMNHITSIAGTPKRPGAYIEGYMFAMFNENQKPNGVEQHF GLFHPNMQPVYPV | |||
| Full Sequence |
|---|
| Protein Sequence Length: 480 Download |
| MEEEWNAEAS SWWCSENEVF PNSREIEDTL QEKMGDQAAQ TQSSQDELEE LLNGDDVKQE 60 EWRIRRHAPM NLYDLFLVIW RSNEIESRCK WEREGRGVVP RWWMAVKSSE NAVKGIEKVV 120 EYRVSTGNEL GSSDQKVMAV LHWIALILSF VATIHNHLGL VEASPDIGVC YGMIGNDLPS 180 ATEVVNLYKT YGIGKMRLFD RNPSALQALR GRQINVTLGI RNEDLPNLAV SQDAVNSWFA 240 SNVEPYLNDI IFNYISVGNE VIPGALGDYV LPVMLSLQNI LDEKNLACIK VTTVVPGSAL 300 GVSYPPSSGE FTSEVSSIMS GIVPFLAQQS SPLMINVYPY FAYASDPANI RLDYAQFTAT 360 SPVVQDGSLS YYNMFDAMVD AFLAAMEKID GVGANVVDVV VSESGWPSDG NGNFTTPGLA 420 ATYNKNYMNH ITSIAGTPKR PGAYIEGYMF AMFNENQKPN GVEQHFGLFH PNMQPVYPVF 480 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| COG5309 | COG5309 | 3.0e-6 | 163 | 469 | 332 | + Exo-beta-1,3-glucanase [Carbohydrate transport and metabolism] | ||
| pfam00332 | Glyco_hydro_17 | 4.0e-116 | 167 | 479 | 315 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAM62473.1 | 0 | 141 | 480 | 10 | 345 | beta-1,3-glucanase bg4 [Arabidopsis thaliana] |
| RefSeq | NP_197533.1 | 0 | 141 | 480 | 10 | 345 | BETAG4; catalytic/ cation binding / hydrolase, hydrolyzing O-glycosyl compounds [Arabidopsis thaliana] |
| RefSeq | NP_197534.1 | 0 | 156 | 480 | 32 | 354 | BG5 (beta-1,3-glucanase 5); glucan 1,3-beta-glucosidase/ hydrolase, hydrolyzing O-glycosyl compounds [Arabidopsis thaliana] |
| RefSeq | NP_197539.1 | 0 | 142 | 480 | 4 | 344 | beta-1,3-glucanase, putative [Arabidopsis thaliana] |
| RefSeq | XP_002302261.1 | 0 | 167 | 478 | 7 | 315 | predicted protein [Populus trichocarpa] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 2cyg_A | 0 | 167 | 479 | 1 | 310 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1ghr_A | 0 | 167 | 479 | 1 | 304 | A Chain A, Crystal Structure At 1.45- Resolution Of The Major Allergen Endo-Beta-1,3-Glucanase Of Banana As A Molecular Basis For The Latex-Fruit Syndrome |
| PDB | 1aq0_B | 0 | 167 | 479 | 1 | 304 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| PDB | 1aq0_A | 0 | 167 | 479 | 1 | 304 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| PDB | 3f55_D | 0 | 166 | 472 | 1 | 306 | A Chain A, Barley 1,3-1,4-Beta-Glucanase In Monoclinic Space Group |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| DC899652 | 345 | 144 | 480 | 0 |
| GR710242 | 280 | 175 | 454 | 0 |
| DY278959 | 330 | 158 | 478 | 0 |
| CO899486 | 217 | 166 | 382 | 0 |
| FC919300 | 340 | 149 | 479 | 0 |
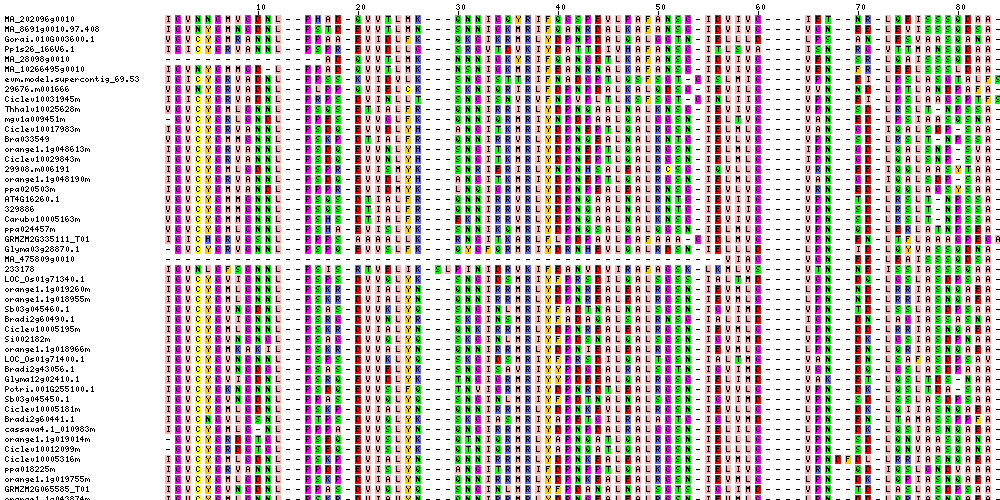
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |