

| Basic Information | |
|---|---|
| Species | Prunus persica |
| Cazyme ID | ppa024759m |
| Family | GH17 |
| Protein Properties | Length: 266 Molecular Weight: 29031.7 Isoelectric Point: 10.091 |
| Chromosome | Chromosome/Scaffold: 7 Start: 8588782 End: 8589763 |
| Description | beta-1,3-glucanase 3 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GH17 | 10 | 255 | 0 |
| IGVCNGMVGDDLPPQAEVVSLYKTNNIPRMRLYDPNPAALEALRGSNIKLLLGPSHRKCMVPKNVRNYANVKFKYIAVGNEVKPSDSFAQFLVPAMQKLQ KAISLAGLANKIKVLTAIDTGVLGGTFPPSIGSFKSEYNALLHPIIRFLVNHQSPLLVNLYPYFAYCGNTQDIRLDYALFTAPSVVVQDGNFGYRNLFDA MLDGVYAALEKAGGGSLKVVISETGWPSAAGTATTIDNARTYRRNR | |||
| Full Sequence |
|---|
| Protein Sequence Length: 266 Download |
| RIIHLTRAPI GVCNGMVGDD LPPQAEVVSL YKTNNIPRMR LYDPNPAALE ALRGSNIKLL 60 LGPSHRKCMV PKNVRNYANV KFKYIAVGNE VKPSDSFAQF LVPAMQKLQK AISLAGLANK 120 IKVLTAIDTG VLGGTFPPSI GSFKSEYNAL LHPIIRFLVN HQSPLLVNLY PYFAYCGNTQ 180 DIRLDYALFT APSVVVQDGN FGYRNLFDAM LDGVYAALEK AGGGSLKVVI SETGWPSAAG 240 TATTIDNART YRRNRKRGLV YLSEK* |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| pfam00332 | Glyco_hydro_17 | 3.0e-112 | 10 | 254 | 258 | + Glycosyl hydrolases family 17. | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
| GO:0005975 | carbohydrate metabolic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| GenBank | AAL30425.1 | 0 | 6 | 254 | 36 | 296 | AF435088_1 beta-1,3-glucanase [Prunus persica] |
| GenBank | AAL30426.1 | 0 | 3 | 254 | 28 | 289 | AF435089_1 beta-1,3-glucanase [Prunus persica] |
| GenBank | ABM74067.1 | 0 | 8 | 254 | 41 | 299 | beta-1,3-glucanase 1 [Prunus avium] |
| GenBank | ACD45060.1 | 0 | 3 | 254 | 28 | 291 | beta-1,3-glucanase [Vitis riparia] |
| Swiss-Prot | P52408 | 0 | 6 | 254 | 36 | 296 | E13B_PRUPE RecName: Full=Glucan endo-1,3-beta-glucosidase, basic isoform; AltName: Full=(1- |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3f55_D | 0 | 10 | 254 | 2 | 262 | A Chain A, Arabidopsis Thaliana Peroxidase N |
| PDB | 3f55_C | 0 | 10 | 254 | 2 | 262 | A Chain A, Arabidopsis Thaliana Peroxidase N |
| PDB | 3f55_B | 0 | 10 | 254 | 2 | 262 | A Chain A, Arabidopsis Thaliana Peroxidase N |
| PDB | 3f55_A | 0 | 10 | 254 | 2 | 262 | A Chain A, Arabidopsis Thaliana Peroxidase N |
| PDB | 3em5_D | 0 | 10 | 254 | 2 | 262 | A Chain A, Crystal Structure Of A Native Endo Beta-1,3-Glucanase (Hev B 2), A Major Allergen From Hevea Brasiliensis |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
 | ||||
| Hit | Length | Start | End | EValue |
| FQ339851 | 228 | 39 | 254 | 0 |
| FN725985 | 242 | 26 | 254 | 0 |
| FC879777 | 267 | 3 | 254 | 0 |
| DY269213 | 267 | 3 | 254 | 0 |
| FC887302 | 267 | 3 | 254 | 0 |
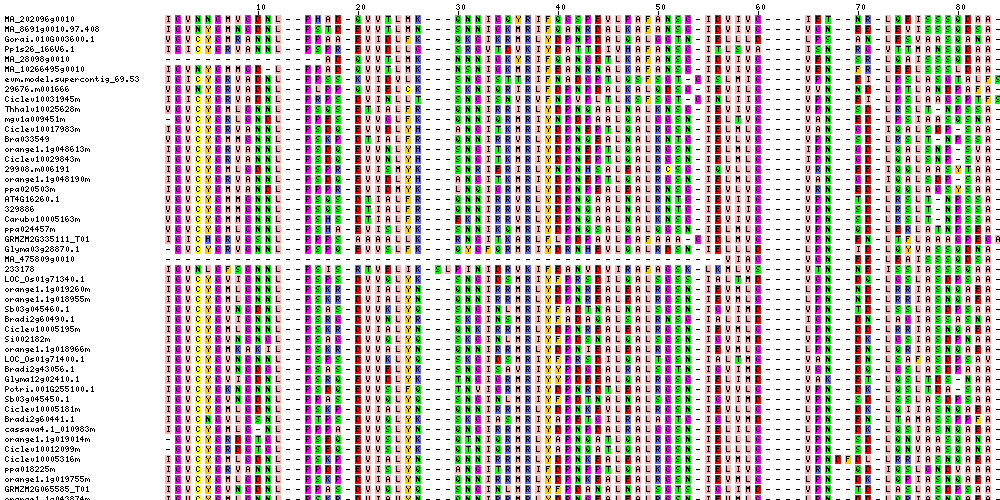
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |