

| Basic Information | |
|---|---|
| Species | Carica papaya |
| Cazyme ID | evm.model.supercontig_17.97 |
| Family | GT5 |
| Protein Properties | Length: 959 Molecular Weight: 108918 Isoelectric Point: 5.2964 |
| Chromosome | Chromosome/Scaffold: 17 Start: 1211641 End: 1227343 |
| Description | starch synthase 4 |
| View CDS | |
| External Links |
|---|
| NCBI Taxonomy |
| CAZyDB |
| Signature Domain Download full data set without filtering | |||
|---|---|---|---|
 | |||
| Family | Start | End | Evalue |
| GT5 | 532 | 764 | 0 |
| VVHIAAEMAPVAKVGGLGDVVTGLGKALQKRGHLLEIIIPKYDCVQYDRIRDLRALDMVVESYFDGRLFQNKIWVGTVEGLPVYFIEPQHPDKFFWRGQF YGENDDFRRFSYFSRAALELLLQAGKKPDIIHCHDWQTAFVAPLYWDLYVSKGLNSARICFTCHNFEYQGAASATELASCGLDVQQLNRPDRMQDNSAQD RINPVKGAIVFSNIVTTVSPTYAQEVRTAEVGC | |||
| GT5 | 762 | 945 | 0 |
| VGCITRLVPQKGVHLIRHAIYRTLELEGQFVLLGSSPISHIQREFEGIADHFQKHEHIRLILKYDESLSHSIYAASDMFIIPSVFEPCGLTQMIAMRYGS VPIARKTGGLNDSVFDVDDDTIPLQFRNGFTFLTPDEQGLNSALERAFSLYSSKPDSWQELVKKDMEIDFSWATSASQYEELYA | |||
| Full Sequence |
|---|
| Protein Sequence Length: 959 Download |
| MAGRLSTYFL CQGFTGLNCN SKHLNVRLFL PSNRSIPASC KMRQRAFSSQ KRQEVKDTSP 60 LPNPTSADLQ QSCVEDSESE NSSINSAANL NVEIENNNIH TAIDLEHGEE KNFDALSVPE 120 PITLELKEPE LRSHPQGGKR VLMRINLPFV ATAMGATLVM NIDREENQSG IQFEELISMI 180 RNAEKNILLL DRARAHAVED LNKILAEKEA LQGEINVLEM KLAETDARIK VAAQEKIHVE 240 VLEDQLGKLC NELGLPSAHD INLQPLSKEL DLLRIENLSL KTDIQALKTE LSSVKNTDER 300 IVMLEKERSL LESSLKELES KLSDSQEDVS KLSTLKVECK DLWTKVENLQ FLLDKATKQA 360 DQAILVLQQN QELRKKVDKL EESLEEANIY KLSSEKMQQY NELMQQKLKL LEERLERSDE 420 EIHSYVQLYQ ESVKEFQETL DSLKEESRKK ALEGPVDDMP WDFWSRLLLI IDGWLLEKKI 480 STGDATLLRE MVWKRDRRVH DVYHSCKEKN EHEAISAFLR LISSATSPGL YVVHIAAEMA 540 PVAKVGGLGD VVTGLGKALQ KRGHLLEIII PKYDCVQYDR IRDLRALDMV VESYFDGRLF 600 QNKIWVGTVE GLPVYFIEPQ HPDKFFWRGQ FYGENDDFRR FSYFSRAALE LLLQAGKKPD 660 IIHCHDWQTA FVAPLYWDLY VSKGLNSARI CFTCHNFEYQ GAASATELAS CGLDVQQLNR 720 PDRMQDNSAQ DRINPVKGAI VFSNIVTTVS PTYAQEVRTA EVGCITRLVP QKGVHLIRHA 780 IYRTLELEGQ FVLLGSSPIS HIQREFEGIA DHFQKHEHIR LILKYDESLS HSIYAASDMF 840 IIPSVFEPCG LTQMIAMRYG SVPIARKTGG LNDSVFDVDD DTIPLQFRNG FTFLTPDEQG 900 LNSALERAFS LYSSKPDSWQ ELVKKDMEID FSWATSASQY EELYAKSVAR ARAAANRT* 960 |
| Functional Domains Download unfiltered results here | ||||||||
|---|---|---|---|---|---|---|---|---|
 | ||||||||
| Cdd ID | Domain | E-Value | Start | End | Length | Domain Description | ||
| TIGR02095 | glgA | 2.0e-66 | 530 | 764 | 238 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| TIGR02095 | glgA | 3.0e-75 | 762 | 947 | 186 | + glycogen/starch synthase, ADP-glucose type. This family consists of glycogen (or starch) synthases that use ADP-glucose (EC 2.4.1.21), rather than UDP-glucose (EC 2.4.1.11) as in animals, as the glucose donor. This enzyme is found in bacteria and plants. Whether the name given is glycogen synthase or starch synthase depends on context, and therefore on substrate [Energy metabolism, Biosynthesis and degradation of polysaccharides]. | ||
| PLN02316 | PLN02316 | 2.0e-115 | 523 | 944 | 462 | + synthase/transferase | ||
| PLN02939 | PLN02939 | 5.0e-121 | 753 | 957 | 205 | + transferase, transferring glycosyl groups | ||
| PLN02939 | PLN02939 | 0 | 1 | 759 | 764 | + transferase, transferring glycosyl groups | ||
| Gene Ontology | |
|---|---|
| GO Term | Description |
| GO:0009058 | biosynthetic process |
| Annotations - NR Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| EMBL | CAA16796.1 | 0 | 1 | 775 | 1 | 782 | starch synthase-like protein [Arabidopsis thaliana] |
| EMBL | CAA16796.1 | 0 | 754 | 958 | 839 | 1071 | starch synthase-like protein [Arabidopsis thaliana] |
| RefSeq | NP_193558.3 | 0 | 1 | 775 | 1 | 779 | ATSS4; transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | NP_193558.3 | 0 | 754 | 958 | 836 | 1040 | ATSS4; transferase, transferring glycosyl groups [Arabidopsis thaliana] |
| RefSeq | XP_002274716.1 | 0 | 1 | 763 | 1 | 747 | PREDICTED: hypothetical protein [Vitis vinifera] |
| Annotations - PDB Download unfiltered results here | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
| PDB | 3guh_A | 1e-38 | 532 | 946 | 3 | 475 | A Chain A, The Structure Of Endoglucanase From Termite, Nasutitermes Takasagoensis, At Ph 2.5. |
| PDB | 2r4u_A | 1e-38 | 532 | 946 | 3 | 475 | A Chain A, The Structure Of Endoglucanase From Termite, Nasutitermes Takasagoensis, At Ph 2.5. |
| PDB | 2r4t_A | 1e-38 | 532 | 946 | 3 | 475 | A Chain A, The Structure Of Endoglucanase From Termite, Nasutitermes Takasagoensis, At Ph 2.5. |
| PDB | 2qzs_A | 1e-38 | 532 | 946 | 3 | 475 | A Chain A, Crystal Structure Of Wild-Type E.Coli Gs In Complex With Adp And Glucose(Wtgsb) |
| PDB | 3d1j_A | 2e-38 | 532 | 946 | 3 | 475 | A Chain A, Crystal Structure Of E.Coli Gs Mutant Dmgs(C7s;c40 |
| Metabolic Pathways | |||
|---|---|---|---|
| Pathway Name | Reaction | EC | Protein Name |
| starch biosynthesis | GLYCOGENSYN-RXN | EC-2.4.1.21 | starch synthase |
| EST Download unfiltered results here | ||||
|---|---|---|---|---|
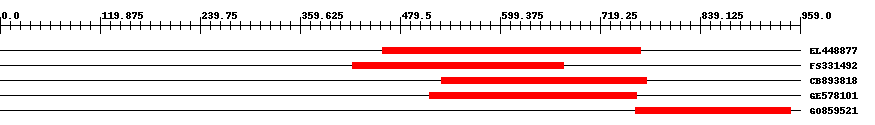 | ||||
| Hit | Length | Start | End | EValue |
| EL448877 | 311 | 458 | 768 | 0 |
| FS331492 | 254 | 422 | 675 | 0 |
| CB893818 | 247 | 529 | 775 | 0 |
| GE578101 | 249 | 515 | 763 | 0 |
| GO859521 | 187 | 762 | 948 | 0 |
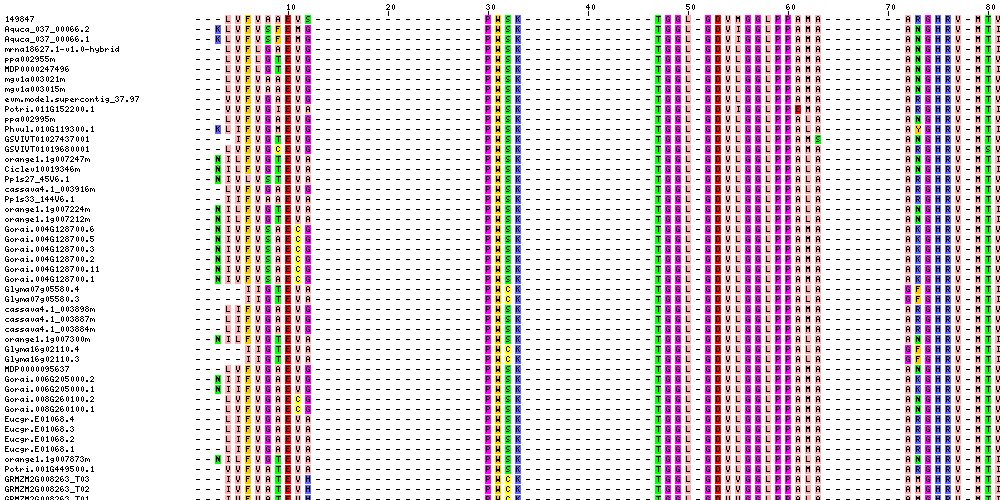
| Sequence Alignments (This image is cropped. Click for full image.) |
|---|
 |